hAT-5_Mam
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001335 |
|---|---|
| TE superfamily | Tag1 |
| TE class | DNA |
| Species | Mammalia |
| Length | 3540 |
| Kimura value | 32.08 |
| Tau index | 0.9753 |
| Description | hAT-Tip100 DNA transposon, hAT-5_Mam subfamily |
| Comment | - |
| Sequence |
CAGGGTAGATAAAAATCNATNATTTTTNAAAAAAAAATTAAAAATCGGATTTTTTAATTTAATCGGATTTTNTANTTTTTNTNGTTAAGATGTTTTNTTAAGAAAAAGAAACCTATATAAAGATAGTTTTAATTTAAGATATGTTAAAGCTCAAAGATATCATCATGGAATAGGGATTATAACTTCTAATTATATANTATTTGAGACAATATATTCATGTAACCTTTTTAGAAAAGTTTTATAAATAGGTCACAACAGGCCACGGACTATATTAGGGGCCCAATCTTATGGGGTTCCCAGAGGCTCCTCTATAGATTANTNAGAAAGATTAAAGAAAACCACTCTACCCAATGGGACTCAGTGCTCAGTCGATCTAGAAGATCCCATTAAGCATCTCTGTGATGCTTAGTTTCAGTTCTCAAACTGTGGATTTGTGTCTCCAGAGATAACATCCTTGTTAACAGCAGTATATGGAGATGAGAGATAACAGACCTGATCAGCCCTATTGTCTCTCTGCAAGTTTGTGTACACAGAGTCAAGCCCTTACCTCTCTCTAAAAGTGCAAAGTTTCAAGAAGTTAAATGAATAGAGGATTGTTGGGGGTGGAGTAGATCTGGGCAAGGAGGAGGAGTCTGGAGATAAATGCAAGGAGGGAGGGGCAGTAGAAACAAAAGAGAAACATTTTCGAGCAGAATATTCCAGAAGTCTTGAGGGATAAGTTTCTGAGTATAGCTTCCATTGATTTGAGATCTAGTGAGGGAATGGTATGGTAGAAGGGAAAACCTATCATGGCAGCAGGCCGTAAAAGAGACCCGTTTGGGAATATTTTAATGAAGTTCCTCTNCCTGTGGGTAAGATAGGCATGCGTGCAAAATGCAAACAGTGCAACAAAGAAATGCAAGGCCTGGTTGCCCGAATGAAACAGCATCGTGAGAAGTGCTCCTTCTCAGGAGGAAGCTGCTTTGAAGATGATGAATGGAACATGTCTGAACATGCGGNATNTCTAGGTTAGTAAACTTTCTTATTNCATTCTTTTTACTCAAGGACTGCCTGNCTTCTCTGTTGACATCTTTTGGACGGANTNCTAGATTACTTGAATTCTCATAAAAATGTTTGAGNAAAAGCATANNTGTTATTTTAAGGANCTATCATATTCNNAATGTNATNGTGANNTGAAANGATGACTGAAATAAGCAGACCTTCCTTTTGCAGTTTCACTTCNAAAGTAGTNCTGGNCNCNATGAGCNANNCTAAATGNNCAGTNNNNTAGCGACTNCATTAACTNCTTTTGTTCAGGAGGATCNANCCTGAACATACAGGATTCTGACTGTTCATCTTCAGCATTATCACTGTCCACAGTNTCGAANNTATCTGCTAACGATACNAGTGTTCGAATCNAATGCAAATCGCATATTATACCNCCTGGGGNNNGNTGGGATTGAAATCTCCGTTATCCAGAAACTACCATAGATAAGTTTGCAATAAGAGCCAGCAGATTNCAGCGAGNGGTAATTGATGAAAAAATTGCTCGGTTTGTTTATGCAACAAACTCTCCTTTCCGTATGGTTGAGAACACACGCTTCACTGACGTGGTTCAGTCATTGCGATCAGGATACAGTCCACCCGACAGAGCAGATATAGCAGGCAAATTGCTGGANAAAGTGTAAGAGNAAGAAATTNAACGGGGTNCANAANGCCTAGAGGGAAAAATTGTGAACCTGAGGCNNGATAGGTGGAGCAATGNCCGCAGTGGTCCAATAATATGNGCTTGTGTGACAACAGAANAAGGGNCTNTCTNCCTTNCAGAAACAATCGATACATCAGGAAATGCACGCANAGCAGAATATTTNCAAGAAATAGCAGTAAAAGCTGTAACAAACTGCGAAAAAAAATTCAGATGTCTANTACGCAGCTTCGTCACAGACAACGCTGCAAATGTATCAAAGGTNAGAAGAAATCTAGAANAAAGTGAAGAAAGCCCCAAGCTAATAACATATGGCTGCAGCGCTCATTTGATGCACCTCTTAGCCAAAGACTTCGGTGCTCCAGAAATAAAGGCTAATGTTGCNGAAATTGCAAAATACTTCCATAACAACCANTTTGCAGCAGCTGCTCCGAAGAAAGCAGGAGGAACCAAACTAACTCTCCCACAAGACGTGCGATGGAACTCAGTAGCGGATTGTTTTGNGTGATATATTGAGAACTGGCCTAATCTGACGACAATTTGTGAACAAAATCGTGACAAAATAGATGGAACTGTCACAGCCAAAGTTTNCAANATTGGGCTAAAGAGAAATGTAGAACGTGTGCTGACTATCCCGAAGCCTATTTCCGTAGCCTTGAACAAAGTGCAGGGAAATCGCCGTTTTATTGCTGATGCTGTTGAAATTTGGAAGGNACTGAGTGAGACCTTGAAAAGAGAAATGTNCAATGACAGAGTNAAATTACGGGCATTAAAAAAACGAACGGATCAAGCGCTATNTCCAGCTCATTTTCTTGCCAATATTCTCAATACTCGGTACCAGGGTCAATCCTTAACCGCTGAAGAAGAGTTGGCTATGACATGGGCATCCAGCAATCATCCCTCCATAATACCAACTATAATAAATTTCAGGGCTAAGGGCGAACCATTCAAGAAATATATGTTTGCTGATGAGGCTTTAAAGAAAGTCACGCNAGTGAACTGGTGGAAGTCACTTAAGCACCTGGATCCATCANAGACTACTGAAGTGATAATCCAGCTTTTAACAGCAGTAGCTTCTTCTGCAGGCGTAGAAAGAATATTTTCTTCCTTTGGACTAATTCATTCTAAATTGAGAAATCGCTTGGGAACTGAAAAAGCAGGAAAGCTTGTCTTTCTTTTCCAGTCTATGAATAAACAGGAAGAGGAAGGTGAAGACTGAATTNGCTGCGTAACCCAGTATTTTAAGTTTCTCATGTTGACNTGGCTGAAATAATCTATTTAATTTTTGTNAAAANACACATTTCGCGTTTATAANTAAAATNGTTAAAAATTCAAAAATACACGTGCTTGTTTTGTTAAAATAATTATACACGTTTGTTGTTGAAGAAAAATATCCAAAATACATAAGCTTGTTGTTTTAGTTAAATAAAATAATTTAAATGTCTGTTGCCTTTGGTGACGGTTGTTGTCCTCCNAATACAGNATGGCAAAAATATCCCCCAAATATCCATGATTAATCNANTGAATTGGAGATAGTTCACCTCCCAATGACTTCACAAGTATCTGCTTCAATTAGCTTTGGTAAATAAAATAACCAAATCATAATTCAGTTTTCTGATGTAGCTGCAAAATTAATCTGAAAAAATATCAAAATAAATCACTGCTTTAAAATGTATAGTGTGTACCTTCTAAAAATGAAACCTACATCTATCTCTGAGTTGTGAATAATATGTATTAAGGTTATATCAAACAACAAGAATGCACTTTTTGTAGAAAACAATGATTAAATCGAGGCTTCCTGACCAGTGATTTAAATCGTGATTTAAATCGATTTGATTTAAATCAAATCCACCCTG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| hAT-5_Mam | TP73 | 981 | 996 | + | 21.98 | ACATGTCTGAACATGC |
| hAT-5_Mam | TP73 | 981 | 996 | - | 19.94 | GCATGTTCAGACATGT |
| hAT-5_Mam | HSF1 | 692 | 704 | - | 19.00 | TTCTGGAATATTC |
| hAT-5_Mam | HSF2 | 692 | 704 | - | 18.38 | TTCTGGAATATTC |
| hAT-5_Mam | HLH4C | 2103 | 2116 | + | 18.11 | GCAGCAGCTGCTCC |
| hAT-5_Mam | TP53 | 980 | 997 | - | 18.10 | CGCATGTTCAGACATGTT |
| hAT-5_Mam | ERF::FOXO1 | 2875 | 2886 | + | 17.86 | ATAAACAGGAAG |
| hAT-5_Mam | HLH4C | 2103 | 2116 | - | 17.84 | GGAGCAGCTGCTGC |
| hAT-5_Mam | FLI1::FOXI1 | 2876 | 2886 | + | 17.74 | TAAACAGGAAG |
| hAT-5_Mam | ETV5::FOXI1 | 2875 | 2886 | + | 17.35 | ATAAACAGGAAG |
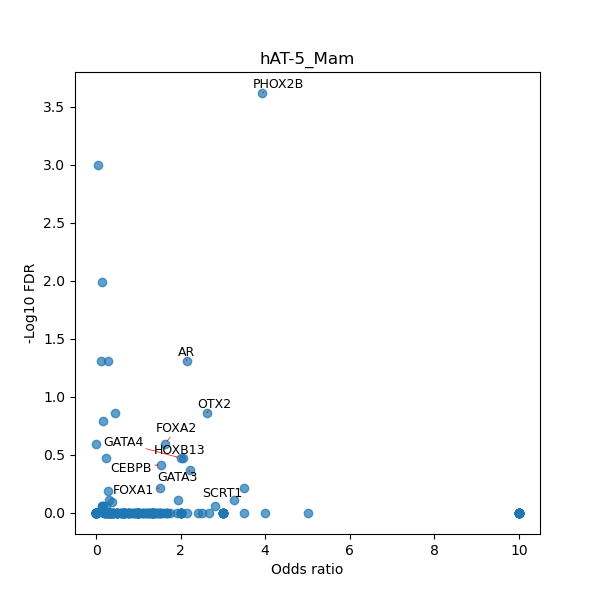
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.