hAT-1_Mam
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001293 |
|---|---|
| TE superfamily | Tag1 |
| TE class | DNA |
| Species | Monotremata,Theria_mammals |
| Length | 3469 |
| Kimura value | 32.43 |
| Tau index | 0.0000 |
| Description | hAT-Tag1 DNA transposon, hAT-1_Mam subfamily |
| Comment | 23 bp TIRs. Near-identical to hAT-1_AMi, which was active in the ancestor of all crocodiles. ORF 1007-2917 encodes a transposase related to other Tag1-group hAT tranposases, like hAT-13_AMi_tp and HAT-25_SM_tp. Occasionally found at orthologous places in marsupial and eutheria. There are copies in platypus, but it is unclear that they are at orthologous sites |
| Sequence |
AAATGTAAAGGCCCGGGAAAAAATCACGGAATAATNNGGGAAAAAAATANNCTTTCCCCTTCTTCTCCGAAGAGTTTTTTGCCCTNTTTCCNNNAAAAATCGGAAAAATTGAAAATTTTTNGAAAAATTAGAAAANTTTGGATTATGAANTNTTTTAGAATGATGAATAAAATGTTTAAGATAAGGTGTTCATATATTGCTACCTGGCCTTTACTCCCCCAAAATGATTTCTAAAAACTATGCAATAGGAAGATTTTTATGTAAAACTATCTACAGTATCAGATCATTACAATTCCTGTACACTGAGGACAAGCAGTACAGTACAACTCTCAAATGCAAGAGCAAGATGCATTTCTGCCTCCAGGGGGCACTGCCATTTTCACAAGAGAAGGAAGAAGGATTCAGTGCCTTCTTTGCTTGTGGGCTTCCTCTTGTTGACTTTTGATCTAGAGGTTGGTGGATATGTTCCTGTCCTCCTGACTAGGCCTCAAAGGACCGGGAGTGGAGTGGACACTGGCAGTAGCATCCATTTATTCCTGTAGAGATTAGAAACAAGGAGCTTATTGCCCTGCTTGCTCAAAAGGCTCAGGACAGAGAGAAGTAAGAGCAAGTCTAAAGGTTTTGGAGTTGGAAGGAAGTTGGAACCTGGCTCCTCCTGGCCCTGTGACCTTGAGATCCACAAAGGAGCTGAAAGAAAGTATACTTCTTTATTGAAAGGTAAGAGTTTAAATTTATCTGAATTCAGAATATTAAACTATATTTTAGCTGCATTAGCATATTTTCATAACCTATGTAAATCATAATTGAATTCCTACTAATCCGATGTTTTGTGGCAAAATCACCTACCCAACCTTAGCAGTGAGGTGAGGCCTGCAGGGCTGGAAAGTTGTTTACAGTTAACTCTGAGGAGACGCCATTACTAAATTTGTACATGCTGTAGCTTAGTGTTTATATGCAAGCTAGCCACTAAACTACATTTTTCCTTTTTTCTCCCTTCAGCAAAATGGTGAGACCAACTTCTAGCATATGGAAGCATTTCCACACTGCAAGTTATTCTGAGGGGGGAAAANACGTGATTTGCAAATACTGTCAAATGAAATATGCATTTCCAAATGCTACTAGGATGACCAAACACATCCTTGTATGTAAGAAGTGTCCTGCAGACATTAAAAAATCCTTCATGGGTATAAGTGAGGAGGACCACGATGAAGAAAGTGCAAGTAGCTCTTTCCAGCGTTCTCGATCCAGAAGTCCTCCTGCTGCTGCTTCTGTAACATCAGCATCATCACAAAGGGTTGCCCAATCAACATCCTCGAGTAACCAAATACTGCAAGCAACTGTATACAAAAAACTCTCAATGGCTTCATTTGTAGACAAGATGACACCTGAAGATCAGGAAAAAATTGACCAAGCTCTTGCAAGAGCAATATATTCTTCTGGAACGCCACTGTCAATAACTGAAAATGCATACTGGCAGGAAGTTTTTAAATTACTAAGACCATCATACCGTTTGCCCAGCAGACATTCTTTGAGCAAGCCTCTTTTAGAGTCAGAATATGAACGGGTAATGGAATCTGTGCAAGGAAGAATCAATGAAGCACTGTGTCTTGCAGTACTTACAGATGGATGGACAAATGTGCGGGGAGAAGGAATCATAAACTTTGTGATAACAACACCTCAACCAGTTTTTTACAGAAGCATTGAAACAGGAGAGAACAGACACACTGCAGAGTATATCAGTTGTAAAATATGTGAAGTCCTACAGAAGACAGGGAGCGGTAAAGTATTTGCTTTGCTGACTGACAATGCAAGTAACATGAAAGCAGCATGGGAGATCATAACGGATAAATANCCACATATTACTGCAATAGGATGTGCATCCCATGGCCTAAACTTGCTTCTTAATGACATCATGAAGTTGGACACTCTCCAAAAGATCTANAGAAGAGCAAAAGAAGTTATAAAACATGTAAAAGCTACTCATATTGTAGCTGCTGTCTTCAAGAAGAAGCAAGAAGAAAAGAATGCAAAGAATAAAATAGTGACACTGAAACTCTGTAGTAAAACTCGATGGGGTGGAGTGGTAATGTCCTTTGAGAGTTTACTAAAAAACAAAGAAGCTCTTCAAGAAACAGTAATAGTTGAGGATCTTAAAGTACCCAGAAGTGTAAGGAACACTGTACTTGATGAGGATGTCTTCTGGGTTCAACTGCAGAACTCCCTGAAGATTCTAAAGCCAATTTCTTCAGCAATCACTGCATCTGAATCAGACAGTGCACTTTTGTCAGACATTCCTTACCAAATGGCCCAGATAAAATCAACTGTTTTTGAGAATCTAAGTGTATTTCCGCTTACAAGTACAGAAGAAAGTAAAGTGAAAGACTTTATTAAGAAACGGGAAGAATTTTGTTGTCAGCCAATTCATGCTGCTGCAAATCTCTTGGATCCAAGATTCAAAGGCAGAAATGTAAGTGATGATAGCACTGCTACTGCATTTGACTGGATTACAAAACAAGCAGCACACTTAGGACTTGATGTTGGAAAGGTACTTTCAAATGTTGCAGAATACAGAACATCTTCAGGTGTTTGGTCCAGGAATGCAGTCTGGGATTCTGCCAAGCATATCCCTCCTGCAACATGGTGGCAAGGCCTTTGCACGACACAGCCTCTGACTCCTCTTGCTTCTCGCCTGCTTCAGATTCCTCCATCTTCTGCAGCATGTGAAAGAAATTGGTCTATGTTTGGAAATACTCGCACCAAATCAAGAAATAAACTAACATGTGAAAGAGTAGAGAAGCTTGTCTCAATCCGGGCAAACCTTCAGTTTTCAGAAGTCCGTGAAGATGATAAAAATGAAGCATCAGAAGAAGCAGAAACTGCTGATAGCGATAGCTCAGAAAATGAGACTGATTAGACAAAGTTCCATGTGGCAAAAGTTGTATTCAATGAGTTTGGGTTTTTTTCCTTTTTTCCCTCAGCTCCTTTTCTGAAAATGAGTGCTTTCTGTATCTGCTTGACACTGTGGAGGACACTAAGTCCAGAAAACATAATAACTTTCTTTGTTTTACTATAATGTTGATGGTTTCTTCTGTTCTCAAGTTTTTGCCTGCTTCTCATAAACTGGCAAAAACTAAGAGATGCTAGATTCCTTAAAGTTCAGCAGCTTGTAGCTCCATTTGTAACAAAAATTAAAAACATTTAAAAGTAAATTCAATTAGTAAATTTGTAATTATTGTCTGAATAAACTATACTTTTAAATTTAATATACCAAACATGATAATTTTATGAGCAAACCAAGTTATACATAATACATTTTTATGTTCCTAAACTAACAAAGTGACTTTAGATCTGTAAAGGGCGCTAAAAAAATAAGTATTGCATATATTTCAATTTTCCCCAAAATTTCCATTTTTTCCCGAAAAACGTTTTCTCCCATGGCTTCAAAATTTCCGGAAATTTTACATCTCTA
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| hAT-1_Mam | WRKY29 | 434 | 442 | - | 15.92 | AAAGTCAAC |
| hAT-1_Mam | PI | 978 | 991 | - | 15.89 | AGAAAAAAGGAAAA |
| hAT-1_Mam | WRKY11 | 433 | 443 | + | 15.89 | TGTTGACTTTT |
| hAT-1_Mam | Esrrg | 665 | 673 | - | 15.85 | TCAAGGTCA |
| hAT-1_Mam | WRKY3 | 433 | 443 | - | 15.85 | AAAAGTCAACA |
| hAT-1_Mam | NR5A1 | 665 | 676 | - | 15.84 | ATCTCAAGGTCA |
| hAT-1_Mam | CTCFL | 362 | 369 | + | 15.72 | CAGGGGGC |
| hAT-1_Mam | DOF1.5 | 2960 | 2975 | - | 15.68 | AGGGAAAAAAGGAAAA |
| hAT-1_Mam | ZNF680 | 1580 | 1590 | + | 15.53 | CAAGGAAGAAT |
| hAT-1_Mam | TCX6 | 3252 | 3266 | + | 15.51 | TTTTAAATTTAATAT |
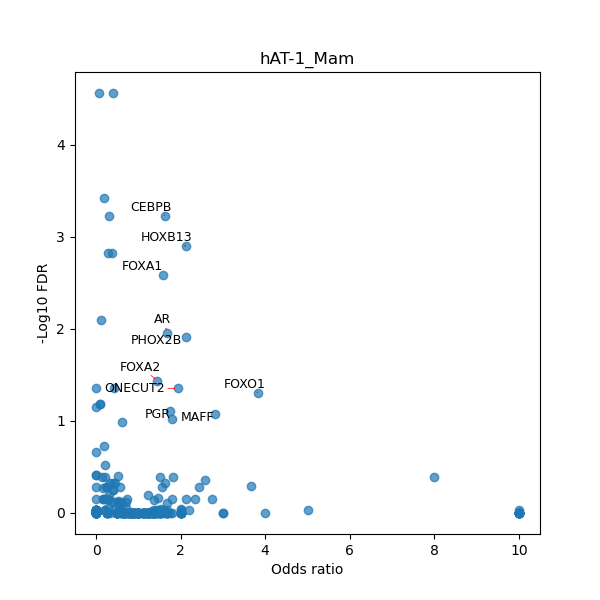
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.