Zaphod4a
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001290 |
|---|---|
| TE superfamily | Tip100 |
| TE class | DNA |
| Species | Eutheria |
| Length | 1823 |
| Kimura value | 29.46 |
| Tau index | 0.0000 |
| Description | hAT-Tip100 DNA transposon, Zaphod4a subfamily |
| Comment | 8 bp TSDs. Only 7 bp TIRs. Non-autonomous. Remaining ORF 1022-1567 encodes the C-terminal quarter of a transposase closest to Arthur1_Cin_tp and Arthur3_Api_tp, but closer to original Zaphod than to original Arthur. Sloppy hairpin structure at 637-736. Pos 303-338 is inverse duplicated at pos 576-541. Present at orthologous sites in elephant and humans, absent in opossum. |
| Sequence |
CAGTGGCGAAACTAAGCCGATGCAGCCGATGCGTTGCATCGGGGCCCATGGGCTATAAGGGGCCCATGTGAGTTGAAAGTTTCAATAGTATTGATCGTGGCCACTGAATAGGCTAATGCATTTAAAAATCAGTATGACATCATTAGCAAACAAACGAAATTGTACTTGAATAGAATTCAACTTTGGCTGGATTTTCTTAAAAATCTGGCTACAGTATTATGGCGCTGGCGCACATATTTTAATTTGGCGCTGGGCCTCCATGACAATAGAGTAGGTGAAATTCCATAAATGTTCGAGCGCATCTATACTCTATTCCAATGACCTACGGAGTAATAAATCATTCTGTCAGTTGCGCAGTAACATTTAAGTCCCACCCTCCTAATCCCTAATTGGCTAAGCAGTTGCAAGACCAAATTACATAGTACTGAAGAAAATATAAACACGAGCATGTAAACAATATCACAATAGATCACTGTACTTGTCAAATTTACCCTGCGTAATTCAGACTTTCTCTATTTCGCCAAAGAACGCCTCTCGCGCATTTATTACTCCGCAGGTCATTGGAATAGAGTATAGCACGCCTGGATTATTATAATGTGCAAAGAGATCGTACATATATCTCGCCGGTACGTCGGGAGCCCCTATGAAAGCTCTTTGATCTTTGCCGGCTTTTTAGCCGAAAGGATTCGGAAGCCTTTGTCGGCATCGGCAAAGACAAAGAGCTGTCGCAGCGGTTGATAGCCCTTTGATATTTGGCCGCTTTTTAGCCGAAAGGATGCGGAAGTCTTTGTCGGAATCATGTATTAATATATAGTGGACAATGCCATTGTCTTAAAGGCTAGGTGAAAATTGATTTTTTAATCATGTTTCCCGCATCGAAAAACTCCCTTGGCCGAAATTTCGCCGAGATCCGACGATATTTAAAANATTTTTGCATCGTCTGCTTCGGCCTGTCGATATGCCGAGAGCCAATATGATAGCTCTCTGATCTTCGGCAGTTTTTTAGGCGAAGGGATTATAGTTCATCATTTTTCATCGTGTAATTGATATAGTAACTACCCAGCTAGACACGAGATTTCAAGGGCTAAGTACAGTTGCTGACTTGTTTGCATTTCTTACTCCAACCCAATTAATTCAAATGGATGAAAAGTCCATTGTGAAAAATGCACAGAAGGTGCAAAAGAAATTTTCAAGTGACCTATCAGAATCATTACCAATGCAATTACTGCTTTGCGTAAAATCATTAAAAAGTGAAATTGAAAAAATTAAAACTATCTGCGAATTTGCCGAGATAGTTATTATAAAATACTACGACATGAGTGCAAGTTTTTCTGAAGTGTGTTCTTTATTAATTTTATTTTTGACAATTCCCGTCACTGTATCTATAGCAGAACGTTCTTTCTCCAAATTAAAACTAATTAAAAACTATTTAAGGAACACTATGGGACAACAACGTTTAACAGAGTTAGCACTACTCAGCATAGAAGCTCAAGCAGCTGAAAAAATGGAAGTTGCAAAACTTATCGATGATTTTGCTAATGCCAAATCACGACGAAGACCATTCTAAGTTTGCTTAAAATTTTTAAATTTGTTTATATTGACAGAATATTGTCTTTGTTTATTATATTGTCATGTTTTGCTTTAGATTTTAAGTTTCAAAGTTTGTTTATATAGTAATTTTTTTGTTTTATCAAAAAGAATGCTATTTGGTTATTAATAAAGTTATGTACCTATGTATGGCAAAATAAGCTCTTTTTGTTGTACGTAGGGGAGGCCCACATTAGACTTTGCATCGGGGCCCATTTTTACTAGTTATGCCACTG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Zaphod4a | JUND | 134 | 144 | - | 17.13 | AATGATGTCAT |
| Zaphod4a | JUN | 134 | 143 | - | 17.08 | ATGATGTCAT |
| Zaphod4a | Sox21a | 817 | 831 | - | 17.07 | GACAATGGCATTGTC |
| Zaphod4a | CG4360 | 1750 | 1762 | - | 16.89 | ACAACAAAAAGAG |
| Zaphod4a | SoxN | 819 | 829 | + | 16.87 | CAATGCCATTG |
| Zaphod4a | TGA6 | 134 | 143 | - | 16.80 | ATGATGTCAT |
| Zaphod4a | FOXE1 | 1663 | 1674 | - | 16.52 | CTATATAAACAA |
| Zaphod4a | TCF7L1 | 743 | 754 | - | 16.36 | AAATATCAAAGG |
| Zaphod4a | ATF2 | 134 | 143 | - | 16.33 | ATGATGTCAT |
| Zaphod4a | Gsx | 1411 | 1430 | - | 15.90 | AATAGTTTTTAATTAGTTTT |
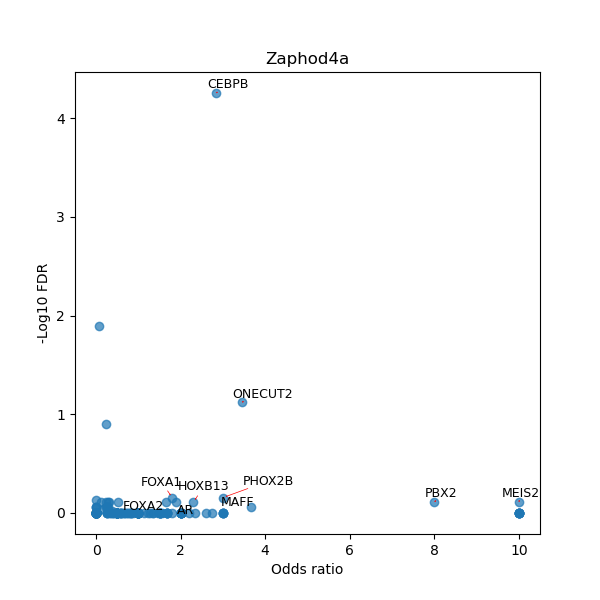
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.