Zaphod3
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001125 |
|---|---|
| TE superfamily | Tip100 |
| TE class | DNA |
| Species | Eutheria |
| Length | 2624 |
| Kimura value | 22.08 |
| Tau index | 1.0000 |
| Description | hAT-Tip100 DNA transposon, Zaphod3 subfamily |
| Comment | Zaphod3 encodes an ORF (~821..2293) that is roughly equally similar to that of Zaphod and Arthur. MER97a-d are non-autonomous, internal deletions of Zaphod3. |
| Sequence |
CAGTGGCGTACCAAGGGCGGGGCGGTGGGAGCGGTCCGCCCCGGGTGCAGGCAATAAGGGGGTGCATTGTCTGTAGAGAATTTAAAAACAATAATAAAACCGACTAAAAGTCGGTCTGCTTTTTATTATCACCATGCGCCGGCAATTCTAAACAATGTCAGTGATAAAATACTCCTCCCCGAAAAATCTTTTGTTGGTCTAAGTTCTAAACAATTGCTGCGGTTACTGTTGAGTTTTAATAATATATATGTAAGCTTCAAATTAGCACATTTTTATTACTTATCCTTTAATAAACATTGTATTCTACATGGAAGTTAATTCGGAGAACTCCCAGTTATACAGTCGGCCCCCGACACACGCGGACTCAGCTACACGCGTTCGTTTCGAGAGTAAGTTCGTAACGGTTCGGAATCGTTCGAGCTCGCTTCGGGCGCAGTTCGTGTCTCCAACCCCTGTGGTACTACATATTCCTGCGTTTAAACAGTAGATTCGAAATAAACAATGATAGCACAGTGATTGTAAAGACGAAGAAACAGAACTTGAGTTACTTCAATTCTGTCATTCTATGTGACCACTTGGAGTTTTTATTTGTGTTTAAAATTTAAAACAGTGAAACAGAGTGCGAACTGCGAGGTGTAATATTTTTGTTTGGTAAGTGCAAATTTTAGTTCATACATGAAATATTTTACTGAATTTGAATAATATCTTTAAAATTGAAATTTATTCTTTTTAAATTGTTAATTGTTTTAAAACTAAAGAATGAATCAAGAAAATAAAACATTACATCAGTGGTACGATTTAGTAGTGGCAGATACTGAAAGCANAATGCCTNGGAATTCTTTCTGNTAGCACTCTNGATGCTTCGAACACTGATCAAATGTGCGAAGTGGTCAGATATGNCNATATTGAAGATGGTAAAGTTGAAATAAAGGAATCATTCTTGGGCTTCTTTCCTATTTCCGGGAAGACTGNTGCTGAACTTNCAGAAGAGANNTCGAAACAACTGGACGGTGATGGGCGGGACATAANCNTCTNCTGTGGNCAAGGATATAACGATGCTGCANNTGCGGCCGGTATTCACGGTGGNGTTCANGCAAAAATCGAAGANATTAATCCCAAATCNTTATTTNTATCTTGTGCAAATCATTCTCTGAACCTTTGCGGAGTTCACTCTTTTGGAAGTATTTCTTCATGCGTGACGTTTTTTNGAACTTTGGAAAAAAATTATTCATTCTTTTCAGTCTCACCTCATCGATGGAAAACGCTGCAGAATGTAGGTATAACAGTGAAAAGACTTTCCCAGACAAGATGGAATGCTCANTATGAAGCTGTGCGCGCAGTANAGACAAATTTTGAAAAGTTAATCTCAACCTTTGAAGTACTGTGTAATCCAAAAGAAAATGTGGACNCAAGAGGATCNGCTCAGATTTTNCTCTCTGCTGTATGCGATTTTTCTTTTCTGAGTTATCTTTTTTTCTGGTGTGAAGTTTTAGATGAGGTTAATCAGACACAAAAATATTTGCAAACAGCCAGAATCAGCCTTGAACAATGTACAGTGAAACACCAAGCTTTAAAATTGTTCCTTGAAGATCGGCGCACAGAAATTGTGGAGAAGGCCATTAACTATGCAACAACAAAATGTAAGGAAATGGACATTTACATAGAAAAAAGAATCAAATTTCGAAGAAGAATGCCAGGAGAAACGACAAAAGATGCTGGTCTTACATTGCCAGAAGAAATCAAAAGGGCAATGTTTGAACGCCTCGATCGTTTTCACCAAGAACTGGACACTCGTTCTAAAGCAATGGATCAAATAATGTCAATGTTCGCTATCATTCAGCCATTTTCTCTGATTTTTGCAGAAGAAGAAAAACTTCGGAAGTTTTTACCAAATATAATAGAAATTTATGATGAATTTTCTGGTGAAGATATTTTAGTGGAAATTTTTCGACTGCGGAGACATTTGAAAGCCGCTAGAATCGATCCCGAAGAAACAAAGACATGGACAGTATTGCAATTTCTGGAATTTATTGTGAAATGGGATTTTTATGAATCTCTGCCAAACTTATCCTTATGTTTAAGACTTTTCCTAACTATTTGTATATCTGTTGCTTCATGTGAAAGAAACTTTTCGAAATTAAAATTAATAAAAAGTGTTCTTCGATCAACTATGAGCGAAGATAGATTGACAAATCTGGCTATACTGTCTATTGAACATGAATATGCGAAGAAGATCAATTTTGACGAAGTCATTGACAAATTTGCAGAAGTTAAGGCTCGAAAACAGAAACTGTAATGTTATTATTCATTACTGCGACAGACCAATATGTAGGTATAAGTTTTTTCCTTTTTTCAAAAAATACATTAATGTAATTAAAAAGTATTAATCCATTACTTTTTTTCCTTTTTTGTACTGTAATATTTATTTTTTATTTTTTATACTGGCATGATTATATATACAAAGTTCAATAAAAGAAAATTTTCACTGTCTGCGTTTCTTTTCTGGCCATTATTATTATTCGTTTCATTTCATGATTATTACTGAAAATAATTTTGTCGTATAGAGGAGGGGGGTGTTAAAAAATGATCCGCTCCGGGTGTCAAATACGCTAGGTACGCCACTG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Zaphod3 | NUC | 1700 | 1711 | - | 17.44 | TTTTGTCGTTTC |
| Zaphod3 | DOF3.4 | 2393 | 2409 | + | 17.42 | TACTTTTTTTCCTTTTT |
| Zaphod3 | IDD11 | 1700 | 1711 | + | 17.34 | GAAACGACAAAA |
| Zaphod3 | DOF5.1 | 2391 | 2409 | - | 17.29 | AAAAAGGAAAAAAAGTAAT |
| Zaphod3 | MYB118 | 395 | 406 | - | 17.23 | AACCGTTACGAA |
| Zaphod3 | CDF5 | 2391 | 2411 | + | 17.03 | ATTACTTTTTTTCCTTTTTTG |
| Zaphod3 | IDD1 | 2552 | 2563 | + | 16.82 | TTTTGTCGTATA |
| Zaphod3 | IDD7 | 1698 | 1711 | + | 16.80 | GAGAAACGACAAAA |
| Zaphod3 | MGP | 1700 | 1712 | + | 16.79 | GAAACGACAAAAG |
| Zaphod3 | IDD11 | 2552 | 2563 | - | 16.64 | TATACGACAAAA |
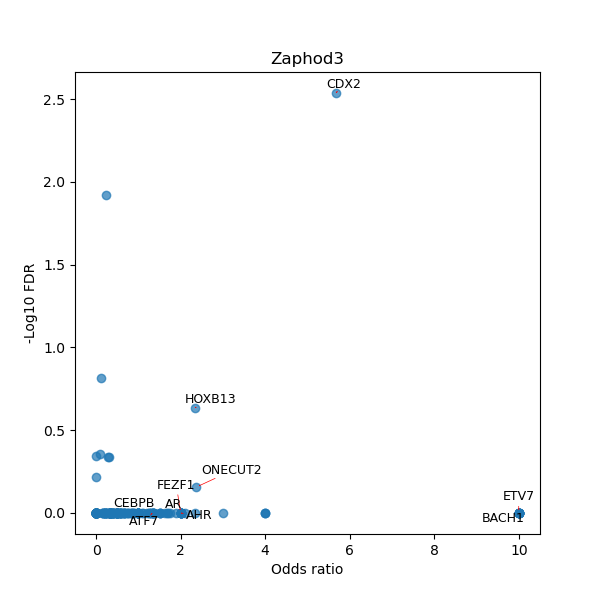
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.