L2
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000358 |
|---|---|
| TE superfamily | L2 |
| TE class | LINE |
| Species | Mammalia |
| Length | 3082 |
| Kimura value | 37.63 |
| Tau index | 0.7930 |
| Description | L2 (LINE2) retrotransposon |
| Comment | L2 (LINE2) spread before the mammalian radiation, and copies are only 65-75% similar to the consensus. The 5' end is probably incomplete. The 3' end is also incomplete, but this terminus is extended by three model variants: L2a_3end, L2b_3end, and L2c_3end. Note that, whereas L1 is A- and purine-rich in the coding strand, L2 is C- and pyrimidine-rich. |
| Sequence |
ACCTCCNAGCCTCCATCGGCAGGGGTCAAGAGGGAGCCCCATGCGCTCNGAGTAAGACCGCCCTCCCCNGGCCTCCACGCCNCCATCGGCAGGAGTNAAGAGAGGGCCCCAGAACTCTGCGTGCTCCTGGCAAGACTCCNACTCCCCACTTCCCCCTCAACACTCACAGCCTCGCCACCACTATCTGCCTTGAGACCATCAGGACTAATCTCATCATGCCCTTTATCCCTGTTTACGTCAGTCACCACAAGAAGCCCAAGACTCCCAGGTCATCCCTCATCCCCCGGACTCTNATCCCCATACGGCTCTGTACATCTCCTNTTCTGACCCCATCTCCCACCCCTCCCCAACTCCTGAAACCCTTCCACTGTGCCCTCTGGAACTCACGGTCCGTCATCAGCAAAATCCCCCGTATCCTCAACCTCTTCTCTGAACGTTCCCTTCACCTTCTTGCTCTAACNGAAACCTGGCTCTCCCCTGAGGACACTGCTTCCCCTGCAGCCCTCTCAAGTGGTGGCTGTTTTCTCTCCCACANCCCTCGTACCACTGGGCCTGGAGGTGGGGTAGGTGTCCTCCTTGCTCCTCATTGCCGCTTCCAGACCATTCTCCCTCCCTCCTCCCTAAAACACCCCAGCTTTGAATCTCATGTCATCAGACTATACCACCCGCTACCCCTCCTTGTTGCAGTCATCTACCGACCTCCGGGTCACTCCCCCTCATTCCTTGAAGATTTTAGCTCCTGGCTCACTGTCACTCTCTCCAACACTACTCCTGTCATAATTCTTGGTGATTTCAATATCCACGTAGATGATCCTTCCAATACCCTGGCCTCTCAGTTCCTTGACCTCCTCTCCTCCAATGATCTTGTCCTCCACCCTACCTCAGCCACTCACTCCCATGGTCATACCCTAGACCTTGTCATTACCAATAACTGCAACCCCTCCATAATCTCAATTTCAAGCATCCCACTCTCTGACCACCACCTCCTATCTTTCCAGCTCACTCCCTCTAGTACCCCGACTCCAACAATTCTTCGACCCCACCGGGACCTCCAATCCATTGATCCTACCACCTTTTCACTGTCCCTCACCCCCCTCATGTCCTCACTTCCCTCCTTACCCAGCTTAGATTCCATGGTCNATCATTATAATCACTCCCTTGCATATACCCTCAACTCCCTTGCCCCTCTCTCGCTTCGTCGTACTCGCCTGGCAAAACCNCAACCCTGGTTAAATCCAACTCTCCGCCTACTCCGCGCCTGCACCCGNGCAGCTGAACGTGGCTGGAGAAAAACACACAACCATGCTGACTGGTCTCACTTTAAATTCATGACCACNAACCTCAAGTGGGCCCTTAATGCTGCCCGGCAATCNTACTACATTTCCCTAGTCCATTCACTCTCCCACTCTCCTAGACGACTATTTCACACCTTCTCCTCTCTCCTCAAACCTCCAACACCTCCTCCCCTATCCTCACTCTCAGCTGATGACCTTGCTTCCTATTTCACTGAGAAAATAGAAGCAATCAGAAGAGAACTTCCACANNCTCCCACCACCACATCTACCCACCTACCTGCATCTGTGCCCACATACTCTGCCTTCCCTCCTGTTACTACGGATGAACTGTCCGTGCTCCTATCTAAGGCCAACCCCTCCACTTGTGCACTAGATCCCATCCCCTCTCGCCTACTCAAGGACATCGCTCCAGCAATTCTCCCCTCTCTCTCCTGCATCATCAATTTTTCCCTCTCTACTGGATCATTCCCATCAGCATACAAACATGCTGTNATTTCTCCCATCTTTAAAAAACAAAAATTCTCCCTTGACCCCACNTCCCCCTCCAGCTACCGCCCCATTTCTCTGCTCCCCTTTACAGCAAAACTCCTCGAAAGAGTTGTCTATACTCGCTGTCTCCAATTCCTCTCCTCCCATTCTCTCTTGAACCCACTCCAATCAGGCTTTCGTCCCCACCACTCCACCGAAACTGCTCTTGTCAAGGTCACCAATGACCTCCACGTTGCTAAATCCAATGGTCAATTCTCAGTCCTCATCTTACTTGACCTNTCAGCAGCATTTGACACAGTTGATCACTCCCTCCTTCTTGAAACACTTTCTTCACTTGGCTTCCAGGACACCACACTCTCCTGGTTTTCCTCCTACCTCACTGGCCGCTCCTTCTCAGTCTCCTTTGCTGGTTCCTCCTCATCTCCCCGACCTCTNAACGTTGGAGTGCCCCAGGGCTCAGTCCTTGGACCTCTTCTCTTCTCTATCTACACTCACTCCCTNGGTGATCTCATCCAGTCTCATGGCTTTAAATACCATCTATATGCTGATGACTCCCAAATTTATATCTCCAGCCCGGACCTCTCCCCTGAACTCCAGACTCGTATATCCAACTGCCTACTCGACATCTCCACTTGGATGTCTAATAGGCATCTCAAACTTAACATGTCCAAAACTGAACTCCTGATCTTCCCCCTCAAACCTGCTCCTCCCGCAGTCTTCCCCATCTCAGTNAATGGCAACTCCATCCTTCCAGTTGCTCAGGCCAAAAACCTTGGAGTCATCCTTGACTCCTCTCTTTCTCTCACACCCCACATCCAATCCATCAGCAAATCCTGTCGGCTCTACCTTCAAAATATATCCAGAATCCGACCACTTCTCACCACCTCCACTGCCACCACCCTGGTCCAAGCCACCATCATCTCTCGCCTGGATTACTGCAATAGCCTCCTAACTGGTCTCCCTGCTTCCACCCTTGCCCCCCTNCAGTCTATTCTCAACACAGCAGCCAGAGTGATCCTTTTAAAACGTAAGTCAGATCATGTCACTCCTCTGCTCAAAACCCTCCAATGGCTTCCCATCTCACTCAGAGTAAAAGCCAAAGTCCTTACAATGGCCTACAAGGCCCTACATGATCTGGTCCCCCGTTACCTCTCTGACCTCATCTCCTACCACTCTCCCCCTCGCTCACTCCGCTCCAGCCACACTGGCCTCCTTGCTGTTCCTCGAACACGCCAGGCACGCTCCCGCCTCAGGGCCTTTGCACTTGCTGTTCCCTCTGCCTGGAACGCTCTTCCCCC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L2 | PRDM9 | 2746 | 2765 | - | 22.39 | GGGGGCAAGGGTGGAAGCAG |
| L2 | Klf15 | 329 | 350 | + | 18.20 | CCCATCTCCCACCCCTCCCCAA |
| L2 | TBX21 | 1423 | 1432 | + | 17.58 | TTCACACCTT |
| L2 | TBX20 | 1422 | 1432 | - | 16.68 | AAGGTGTGAAA |
| L2 | IRF6 | 1977 | 1985 | + | 16.60 | ACCGAAACT |
| L2 | PLAG1 | 2753 | 2766 | - | 16.58 | GGGGGGCAAGGGTG |
| L2 | RAMOSA1 | 1715 | 1728 | - | 16.53 | AGGAGAGAGAGGGG |
| L2 | ESRRB | 1993 | 2002 | + | 16.36 | TCAAGGTCAC |
| L2 | TCP23 | 1346 | 1353 | - | 16.36 | GGGCCCAC |
| L2 | ESRRA | 1993 | 2001 | + | 16.33 | TCAAGGTCA |
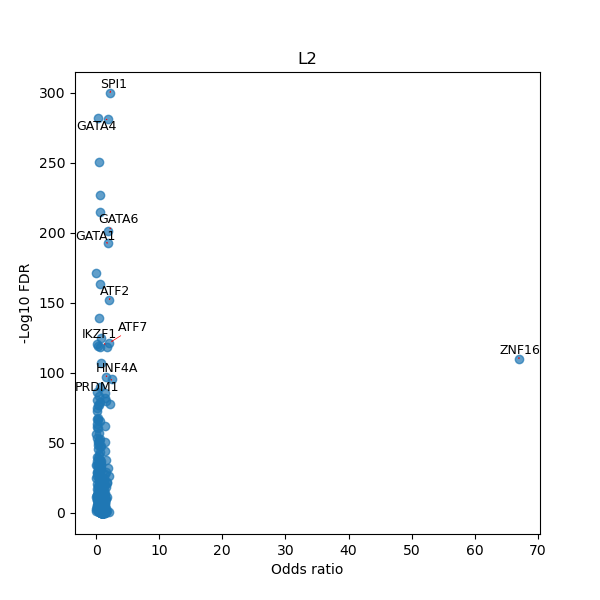
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.