L1P2_5end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000317 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Catarrhini |
| Length | 2275 |
| Kimura value | 5.44 |
| Tau index | 0.8359 |
| Description | 5' end of L1 retrotransposon, L1P2_5end subfamily |
| Comment | - |
| Sequence |
TCCGTTCCAAGATGGCCGAATAGGAACAGCTCCGGTCTGCAGCTCCCAGCGTGATCGACGCAGAAGACGGGTGATTTCTGCATTTCCAACTGAGGTACCTGGTTCATCTCACTGGGACTGGTTGGACAGTGGGTGCAGCCCACGGAGGGCGAGCCGAAGCAGGGCGGGGCATCGCCTCACCCGGGAAGCGCAAGGGGTCGGGGGATTTCCCTTTCCTAGCCAAGGGAAGCCGTGACAGACTGTACCTGGAAAANCGGGACACTCCCGCCCAAATACTGCGCTTTTCCAACGGTCTTAGCAAACGGCACACCAGGAGATTNTATCCCGCGCCTGGCTCGGCGGGTCCCACGCCCACGGAGCCTTGCTCACTGCTAGCGCAGCAGTCTGAGATCGANCTGCGAGGCGGCAGCCTGGCTGGGGGAGGGGCGTCCGCCATTGCTGAGGCTTGAGTAGGTAAACAAAGCGGCCGGGAAGCTCGAACTGGGCGGAGCCCACCGCAGCTCAGCGAGGCCTGCTGCCTCTGTAGACTCCACCTCTGGGGGCAGGGCATAGCTGAACAAAAGGCAGCAGACAACTTCTGCAGACTTAAACGTCCCTGTCTGACAGCTCTGAAGAGAGCAGTGGTTCTCCCAGCACGGCGTTTGAGCTCTGAGAACGGACAGACTGCCTCCTCAAGTGGGTCCCTGACCCCCGTGTAGCCTAACTGGGAGACACCTCCCAGTAGGGGCCGACNGACACCTCATACAGGCGGGTGCCCCTCTGGGACGAAGCTTCCAGAGGAAGGATCAGGCAGCAATATTTGCTGTTCTGCAATATTTGCTGTTCTGCAGCCTCCGCTGGTGATACCCAGGCAAACAGGGTCTGGAGTGGACCTCCAGCAAACTCCAACAGACCTGCAGCTGAGGGNCCTGACTGTTAGAAGGAAAACTAACAAACAGAAAGGAATAGCATCAACATCAACAAAAAGGACATCCACACCAAAACCCCATCTGTAGGTCACCANCATCAAAGACCAAAGGTAGATAAAACCACAAAGATGGGGAGAAACCAGAGCAGAAAAGCTGAAAATTCTAAAAACCAGAGCGCCTCTTCTCCTCCAAAGGATCGCAGCTCCTCGCCAGCAACGGAACAAAGCTGGACGGAGAATGACTTTGACGAGTTGACAGAAGTAGGCTTCAGAAGGTCGGTAATAACAAACTTCTCCGAGCTAAAGGAGGATGTTCGAACCCATCGCAAGGAAGCTAAAAACCTTGAAAAAAGATTAGACGAATGGCTAACTAGAATAAACAGTGTAGAGAAGACCTTAAATGACCTGATGGAGCTGAAAACCACGGCACGAGAACTNCGTGACGCATGCACAAGCTTCAGTAGCCGATTCGATCAAGTGGAAGAAAGGGTATCAGTGATTGAAGATCAAATTAATGAAATAAAGCGAGAAGANAAGNTTAGAGAAAAAAGAGTAAAAAGAAACGAACAAAGCCTCCAAGAAATATGGGACTATGTGAAAAGACCAAATCTACGTTTGATTGGTGTACCTGAAAGTGACGGGGAGAATGGAACCAAGTTGGAAAACACTCTTCAGGATATTATCCAGGAGAACTTCCCCAACCTAGCAAGGCAGGCCAACATTCAAATTCAGGAAATACAGAGAACGCCACAAAGATACTCCTCGAGAAGAGCAACCCCAAGACACATAATTGTCAGATTCACCAAGGTTGAAATGAAGGAAAAAATGTTAAGGGCAGCCAGAGAGAAAGGTCGGGTTACCCACAAAGGGAAGCCCATCAGACTAACAGCGGATCTCTCGGCAGAAACCCTACAAGCCAGAAGAGAGTGGGGGCCAATATTCAACATTCTTAAAGAAAAGAATTTTCAACCCAGAATTTCATATCCAGCCAAACTAAGCTTCATAAGTGAAGGAGAAATAAAATCCTTTACAGACAAGCAAATGCTGAGAGATTTTGTCACCACCAGGCCTGCCTTACAAGAGCTCCTGAAGGAAGCACTAAACATGGAAAGGAACAACCGGTACCAGCCACTGCAAAAACATGCCAAATTGTAAAGACCATCGATGCTAGGAAGAAACTGCATCAACTAACGGGCAAAATAACCAGCTAACATCATAATGACAGGATCAAATTCACACATAACAATATTAACCTTAAATGTAAATGGGCTAAATGCCCCAATTAAAAGACACAGACTGGCAAATTGGATAAAGAGTCAAGACCCATCAGTGTGCTGTATTCAGGAGACCCATCTCACGTGCAGAGAC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1P2_5end | SP1 | 418 | 426 | + | 16.06 | GGGGAGGGG |
| L1P2_5end | KLF9 | 345 | 355 | + | 16.04 | CCCACGCCCAC |
| L1P2_5end | Eip75B | 990 | 999 | + | 16.03 | CTGTAGGTCA |
| L1P2_5end | TCP9 | 339 | 349 | - | 16.00 | GTGGGACCCGC |
| L1P2_5end | KLF16 | 345 | 355 | + | 15.98 | CCCACGCCCAC |
| L1P2_5end | KLF16 | 417 | 427 | - | 15.95 | GCCCCTCCCCC |
| L1P2_5end | IME1 | 335 | 342 | - | 15.91 | CCGCCGAG |
| L1P2_5end | DOF5.1 | 1462 | 1480 | + | 15.89 | AAAAAGAAACGAACAAAGC |
| L1P2_5end | SP4 | 418 | 426 | + | 15.85 | GGGGAGGGG |
| L1P2_5end | ARF10 | 703 | 713 | + | 15.84 | ACTGGGAGACA |
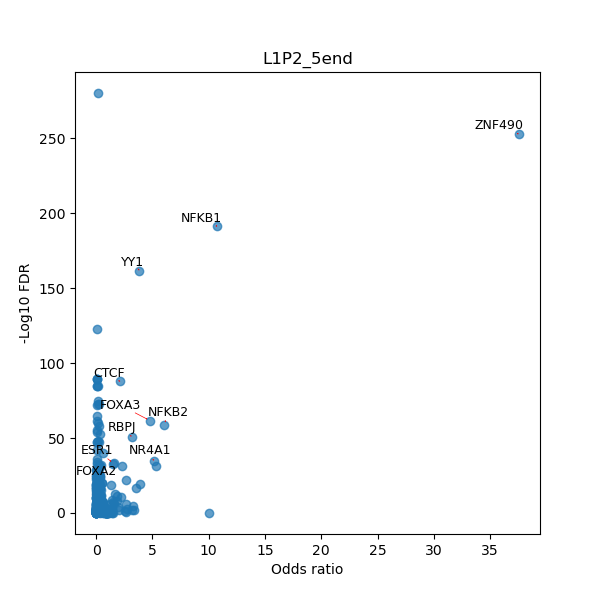
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.