L1P2_5end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000317 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Catarrhini |
| Length | 2275 |
| Kimura value | 5.44 |
| Tau index | 0.8359 |
| Description | 5' end of L1 retrotransposon, L1P2_5end subfamily |
| Comment | - |
| Sequence |
TCCGTTCCAAGATGGCCGAATAGGAACAGCTCCGGTCTGCAGCTCCCAGCGTGATCGACGCAGAAGACGGGTGATTTCTGCATTTCCAACTGAGGTACCTGGTTCATCTCACTGGGACTGGTTGGACAGTGGGTGCAGCCCACGGAGGGCGAGCCGAAGCAGGGCGGGGCATCGCCTCACCCGGGAAGCGCAAGGGGTCGGGGGATTTCCCTTTCCTAGCCAAGGGAAGCCGTGACAGACTGTACCTGGAAAANCGGGACACTCCCGCCCAAATACTGCGCTTTTCCAACGGTCTTAGCAAACGGCACACCAGGAGATTNTATCCCGCGCCTGGCTCGGCGGGTCCCACGCCCACGGAGCCTTGCTCACTGCTAGCGCAGCAGTCTGAGATCGANCTGCGAGGCGGCAGCCTGGCTGGGGGAGGGGCGTCCGCCATTGCTGAGGCTTGAGTAGGTAAACAAAGCGGCCGGGAAGCTCGAACTGGGCGGAGCCCACCGCAGCTCAGCGAGGCCTGCTGCCTCTGTAGACTCCACCTCTGGGGGCAGGGCATAGCTGAACAAAAGGCAGCAGACAACTTCTGCAGACTTAAACGTCCCTGTCTGACAGCTCTGAAGAGAGCAGTGGTTCTCCCAGCACGGCGTTTGAGCTCTGAGAACGGACAGACTGCCTCCTCAAGTGGGTCCCTGACCCCCGTGTAGCCTAACTGGGAGACACCTCCCAGTAGGGGCCGACNGACACCTCATACAGGCGGGTGCCCCTCTGGGACGAAGCTTCCAGAGGAAGGATCAGGCAGCAATATTTGCTGTTCTGCAATATTTGCTGTTCTGCAGCCTCCGCTGGTGATACCCAGGCAAACAGGGTCTGGAGTGGACCTCCAGCAAACTCCAACAGACCTGCAGCTGAGGGNCCTGACTGTTAGAAGGAAAACTAACAAACAGAAAGGAATAGCATCAACATCAACAAAAAGGACATCCACACCAAAACCCCATCTGTAGGTCACCANCATCAAAGACCAAAGGTAGATAAAACCACAAAGATGGGGAGAAACCAGAGCAGAAAAGCTGAAAATTCTAAAAACCAGAGCGCCTCTTCTCCTCCAAAGGATCGCAGCTCCTCGCCAGCAACGGAACAAAGCTGGACGGAGAATGACTTTGACGAGTTGACAGAAGTAGGCTTCAGAAGGTCGGTAATAACAAACTTCTCCGAGCTAAAGGAGGATGTTCGAACCCATCGCAAGGAAGCTAAAAACCTTGAAAAAAGATTAGACGAATGGCTAACTAGAATAAACAGTGTAGAGAAGACCTTAAATGACCTGATGGAGCTGAAAACCACGGCACGAGAACTNCGTGACGCATGCACAAGCTTCAGTAGCCGATTCGATCAAGTGGAAGAAAGGGTATCAGTGATTGAAGATCAAATTAATGAAATAAAGCGAGAAGANAAGNTTAGAGAAAAAAGAGTAAAAAGAAACGAACAAAGCCTCCAAGAAATATGGGACTATGTGAAAAGACCAAATCTACGTTTGATTGGTGTACCTGAAAGTGACGGGGAGAATGGAACCAAGTTGGAAAACACTCTTCAGGATATTATCCAGGAGAACTTCCCCAACCTAGCAAGGCAGGCCAACATTCAAATTCAGGAAATACAGAGAACGCCACAAAGATACTCCTCGAGAAGAGCAACCCCAAGACACATAATTGTCAGATTCACCAAGGTTGAAATGAAGGAAAAAATGTTAAGGGCAGCCAGAGAGAAAGGTCGGGTTACCCACAAAGGGAAGCCCATCAGACTAACAGCGGATCTCTCGGCAGAAACCCTACAAGCCAGAAGAGAGTGGGGGCCAATATTCAACATTCTTAAAGAAAAGAATTTTCAACCCAGAATTTCATATCCAGCCAAACTAAGCTTCATAAGTGAAGGAGAAATAAAATCCTTTACAGACAAGCAAATGCTGAGAGATTTTGTCACCACCAGGCCTGCCTTACAAGAGCTCCTGAAGGAAGCACTAAACATGGAAAGGAACAACCGGTACCAGCCACTGCAAAAACATGCCAAATTGTAAAGACCATCGATGCTAGGAAGAAACTGCATCAACTAACGGGCAAAATAACCAGCTAACATCATAATGACAGGATCAAATTCACACATAACAATATTAACCTTAAATGTAAATGGGCTAAATGCCCCAATTAAAAGACACAGACTGGCAAATTGGATAAAGAGTCAAGACCCATCAGTGTGCTGTATTCAGGAGACCCATCTCACGTGCAGAGAC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1P2_5end | TCP21 | 339 | 349 | - | 16.62 | GTGGGACCCGC |
| L1P2_5end | TCF7L1 | 1008 | 1019 | + | 16.59 | AAAGACCAAAGG |
| L1P2_5end | TCP20 | 339 | 349 | - | 16.43 | GTGGGACCCGC |
| L1P2_5end | TCP15 | 339 | 349 | - | 16.30 | GTGGGACCCGC |
| L1P2_5end | KLF5 | 161 | 170 | - | 16.29 | GCCCCGCCCT |
| L1P2_5end | KLF15 | 162 | 169 | - | 16.27 | CCCCGCCC |
| L1P2_5end | NFkb | 201 | 211 | - | 16.19 | GGGAAATCCCC |
| L1P2_5end | CDF5 | 1451 | 1471 | - | 16.15 | GTTTCTTTTTACTCTTTTTTC |
| L1P2_5end | KLF5 | 418 | 427 | - | 16.10 | GCCCCTCCCC |
| L1P2_5end | MAZ | 419 | 426 | - | 16.08 | CCCCTCCC |
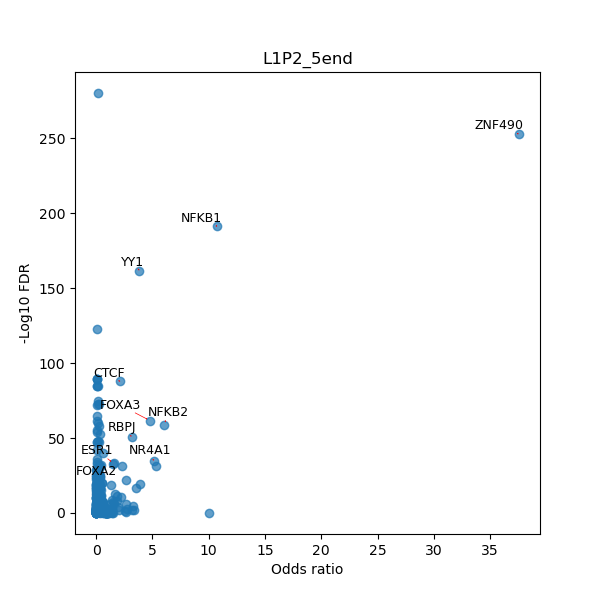
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.