L1MC4a_3end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000279 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Eutheria |
| Length | 2629 |
| Kimura value | 23.95 |
| Tau index | 0.8175 |
| Description | 3' end of L1 retrotransposon, L1MC4a_3end subfamily |
| Comment | - |
| Sequence |
CTAATATCCCTAATATATAAAGAACTCTTAAAANTNGAAAGNAAAAGACCAAAAACCCGATAGAAAAATGGGCAAAAGACATGAACAGACAATTCACAAAAAGANATANAAATGGCCCTTAAACATATGAAAAGATGTTCAACCTCACTCATAATAAGAGAAATGCAAATTAAAACTACACTGAGATACCATTTCTCACCTATCAGATTGGCAAAAATTCAAAAGTTTGACAACACACTCTGTTGGCGAGGCTGTGGGGAAACAGGCACTCTCATACATTGCTGGTGGGAATGCAAANTGGTACAACCCTTNTGGAGGGGAATTTGGCAATATCTAACAAAACTACATATGCATTTACCCTTTGACCCAGCAATCCCACTTCTAGGAATTTACCCTGAAGATACACCTCCAACATGTACAAAATNACATATGCACAAGGTTATTCATTGCAGCATTGTTTGTAATTGCAAAATATTGGAAACAACCTAAATGCCCATNNATAGGAGANTGGTTGAATAAACTATGGTACATCCACACAATGGAGTACTATGCAGCTGTAAAAAAGAATGAGGAAGATCTCTATGAACTGATATGGAGTGATTTCCAGGATATATTGTTAAGTGAAAAAAGCAAAGTGCAAAAGAGTATNTATAGTATGCTACCTTTCGTGTAAGAAAGAAGGGAAAATAAGAAAATATACATGTATCTGCTCATTTGTGCAAAAAGAAACACAGGAAGGATAAACCAGAAACTAATGAGATTGGTTACCTACAGGGGGTGGGTGGGAATGGGGAGAAAGAATGGGGGATGGGAACGGGATGGNAGGGATGAGGAGGGAGTGACACTTCTCTGAGTATACCTTTTTGTATAGTTCTGACTTTTAGAACCATGNTAATGTTTCACATACCTCAAAAATAAATAATTAAAATCAACCAGGATGTGGGGGGAACCCAAAATGGAATACAAACAGNAACAAATGAACCTAACTGTATTNCAAATGAATAACATAACCACACTGAAGGGGATGGGGAAGAAAAGAACTAACCTAAGTAACTTTGGAAAACAGTATTTTGACTGNATACTGTAAGGCTAAAGACAAAAAGAACTGTACACAAATACTGTACTCTAGTTAGTAAATTTGTTTCTCACAGGGGTATGGGTTAGCAATTCTGAAACTACTTTATGTGTATNCTAGGATTGAACAAATAAGTAAATATATTGTAGATAATGAGAGCCAGGTTTCTCACTGTCGGAGAAAGAAGTTACAAATAAGGAAAGGGGGAAGGCTAGAATGAACCCTGTGGTGTTGGATTGGAATCGGAGGTATCAGTATGAACTCATGGTTTTTAACATATATACAGATAGATAGATACAGAAATAAGATATAGATGTATGTGTATGCATGGGTTAGTATACATACATATATTTCCTAGCTCTGTCCGCTGAGAGGGCCTAGAAGCAATGACACCCCAGTAGCAATGAGCACACCTAGCGCCCAGATCTTGGTTTCTAAATACCATTCTCCAATAAAAGGAACCAGGGCTCCTTGGAGAAATGGCTGATTCCAGGGCTGGGGCAGGGAAAATACAAGATGAGCCTGGAACATCTTGTGGTGCCAGAAAGTAAGGAAGTGCTCAAAAAANGATGGGGGCATGTCAAAAGGACACAGGAGCCAACCTGAAGGAGCTCCCAATGGCCAAANCTGGAACAATTTGAGCAACAAAATAAATAATGATAGTANTGGATTATAACCCATAGAATAAAATAAATATCCATGAGTCCATACTGATATAAATAAATAATTGAATAAATAAATAAATGGGGGAGAAGGGACAGCTCTTCCTTACAGNAGAATTCCAATTAATAAATGTAGAAGGAATGANGGAAATAGAAAATCACCATTAGGCAAACACCACAGTAATAATTGTTGCAGGCAAGATCCACCGATGGATGCTAAAATTAGTGGGCGAAAGTTTGAGGAGAAACAGGATATTTGCATAGTCTCAAAGTATCTCCCCCAAGATATTTATTAATTACAAAGGGAAAAATAGTAACTTTACAGTGGAGAAACCCGGCAGACACCACCTTAACCAAGTGATCAAGGTTAACATCACCAGTAATAAGACATATCGACATCATGTACCCCCTGATATGATGCACTGAGAAGGGCACAACATCACTTCTGTGGTATTCTTGCCAAAAATGCATAACCTCAATCTAATCATGAGAAAACATCAGACAAACCCAAATTGAGGGACATTCTACAAAATAACTGACCAGTACTCTTCAAAAGTGTCAAGGTCATGAAAGACAAGGAAAGACTGAGGAACTGTCACAGATTGGAGGAGACTAAGGAGACATGACAACTAAATGCAATGTGGGATCCTGGATTGGATCCTGGAACAGAAAAAAGGACATTAGTGGAAAAACTGGTGAAATTCGAATAAGGTCTGTAGTTTAGTTAATAGTATTGTACCAATGTTAATTTCCTGGTTTTGATAATTGTACTATGGTTATGTAAGATGTTAACATTAGGGGAAGCTGGGTGAAGGGTATACGGGAACTCTCTGTACTATTTTTGCAACTTTTCTGTAAGTCTAAAATTATTTCAAAATAAAAAGTTAAAAAA
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1MC4a_3end | BPC5 | 1267 | 1296 | + | -49.11 | AAATAAGGAAAGGGGGAAGGCTAGAATGAA |
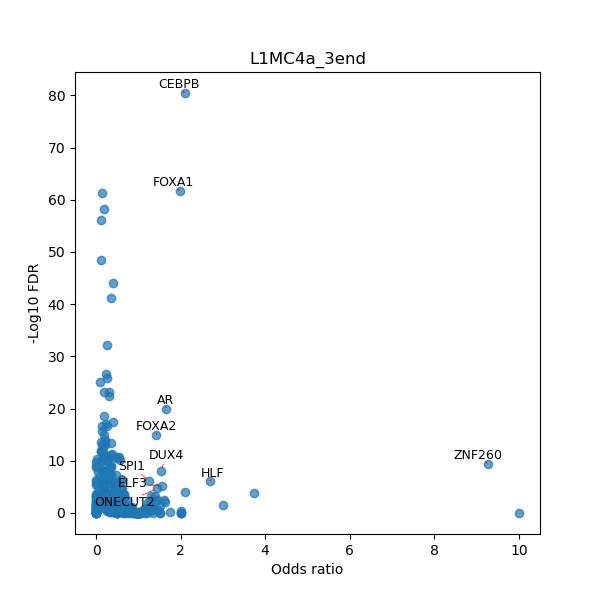
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.