L1MC4a_3end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000279 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Eutheria |
| Length | 2629 |
| Kimura value | 23.95 |
| Tau index | 0.8175 |
| Description | 3' end of L1 retrotransposon, L1MC4a_3end subfamily |
| Comment | - |
| Sequence |
CTAATATCCCTAATATATAAAGAACTCTTAAAANTNGAAAGNAAAAGACCAAAAACCCGATAGAAAAATGGGCAAAAGACATGAACAGACAATTCACAAAAAGANATANAAATGGCCCTTAAACATATGAAAAGATGTTCAACCTCACTCATAATAAGAGAAATGCAAATTAAAACTACACTGAGATACCATTTCTCACCTATCAGATTGGCAAAAATTCAAAAGTTTGACAACACACTCTGTTGGCGAGGCTGTGGGGAAACAGGCACTCTCATACATTGCTGGTGGGAATGCAAANTGGTACAACCCTTNTGGAGGGGAATTTGGCAATATCTAACAAAACTACATATGCATTTACCCTTTGACCCAGCAATCCCACTTCTAGGAATTTACCCTGAAGATACACCTCCAACATGTACAAAATNACATATGCACAAGGTTATTCATTGCAGCATTGTTTGTAATTGCAAAATATTGGAAACAACCTAAATGCCCATNNATAGGAGANTGGTTGAATAAACTATGGTACATCCACACAATGGAGTACTATGCAGCTGTAAAAAAGAATGAGGAAGATCTCTATGAACTGATATGGAGTGATTTCCAGGATATATTGTTAAGTGAAAAAAGCAAAGTGCAAAAGAGTATNTATAGTATGCTACCTTTCGTGTAAGAAAGAAGGGAAAATAAGAAAATATACATGTATCTGCTCATTTGTGCAAAAAGAAACACAGGAAGGATAAACCAGAAACTAATGAGATTGGTTACCTACAGGGGGTGGGTGGGAATGGGGAGAAAGAATGGGGGATGGGAACGGGATGGNAGGGATGAGGAGGGAGTGACACTTCTCTGAGTATACCTTTTTGTATAGTTCTGACTTTTAGAACCATGNTAATGTTTCACATACCTCAAAAATAAATAATTAAAATCAACCAGGATGTGGGGGGAACCCAAAATGGAATACAAACAGNAACAAATGAACCTAACTGTATTNCAAATGAATAACATAACCACACTGAAGGGGATGGGGAAGAAAAGAACTAACCTAAGTAACTTTGGAAAACAGTATTTTGACTGNATACTGTAAGGCTAAAGACAAAAAGAACTGTACACAAATACTGTACTCTAGTTAGTAAATTTGTTTCTCACAGGGGTATGGGTTAGCAATTCTGAAACTACTTTATGTGTATNCTAGGATTGAACAAATAAGTAAATATATTGTAGATAATGAGAGCCAGGTTTCTCACTGTCGGAGAAAGAAGTTACAAATAAGGAAAGGGGGAAGGCTAGAATGAACCCTGTGGTGTTGGATTGGAATCGGAGGTATCAGTATGAACTCATGGTTTTTAACATATATACAGATAGATAGATACAGAAATAAGATATAGATGTATGTGTATGCATGGGTTAGTATACATACATATATTTCCTAGCTCTGTCCGCTGAGAGGGCCTAGAAGCAATGACACCCCAGTAGCAATGAGCACACCTAGCGCCCAGATCTTGGTTTCTAAATACCATTCTCCAATAAAAGGAACCAGGGCTCCTTGGAGAAATGGCTGATTCCAGGGCTGGGGCAGGGAAAATACAAGATGAGCCTGGAACATCTTGTGGTGCCAGAAAGTAAGGAAGTGCTCAAAAAANGATGGGGGCATGTCAAAAGGACACAGGAGCCAACCTGAAGGAGCTCCCAATGGCCAAANCTGGAACAATTTGAGCAACAAAATAAATAATGATAGTANTGGATTATAACCCATAGAATAAAATAAATATCCATGAGTCCATACTGATATAAATAAATAATTGAATAAATAAATAAATGGGGGAGAAGGGACAGCTCTTCCTTACAGNAGAATTCCAATTAATAAATGTAGAAGGAATGANGGAAATAGAAAATCACCATTAGGCAAACACCACAGTAATAATTGTTGCAGGCAAGATCCACCGATGGATGCTAAAATTAGTGGGCGAAAGTTTGAGGAGAAACAGGATATTTGCATAGTCTCAAAGTATCTCCCCCAAGATATTTATTAATTACAAAGGGAAAAATAGTAACTTTACAGTGGAGAAACCCGGCAGACACCACCTTAACCAAGTGATCAAGGTTAACATCACCAGTAATAAGACATATCGACATCATGTACCCCCTGATATGATGCACTGAGAAGGGCACAACATCACTTCTGTGGTATTCTTGCCAAAAATGCATAACCTCAATCTAATCATGAGAAAACATCAGACAAACCCAAATTGAGGGACATTCTACAAAATAACTGACCAGTACTCTTCAAAAGTGTCAAGGTCATGAAAGACAAGGAAAGACTGAGGAACTGTCACAGATTGGAGGAGACTAAGGAGACATGACAACTAAATGCAATGTGGGATCCTGGATTGGATCCTGGAACAGAAAAAAGGACATTAGTGGAAAAACTGGTGAAATTCGAATAAGGTCTGTAGTTTAGTTAATAGTATTGTACCAATGTTAATTTCCTGGTTTTGATAATTGTACTATGGTTATGTAAGATGTTAACATTAGGGGAAGCTGGGTGAAGGGTATACGGGAACTCTCTGTACTATTTTTGCAACTTTTCTGTAAGTCTAAAATTATTTCAAAATAAAAAGTTAAAAAA
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1MC4a_3end | ZNF8 | 518 | 537 | - | 26.91 | GTGTGGATGTACCATAGTTT |
| L1MC4a_3end | ZNF558 | 67 | 95 | + | 22.70 | AATGGGCAAAAGACATGAACAGACAATTC |
| L1MC4a_3end | KLF17 | 773 | 786 | - | 18.55 | CCCACCCACCCCCT |
| L1MC4a_3end | RAX3 | 774 | 785 | + | 18.33 | GGGGGTGGGTGG |
| L1MC4a_3end | ESRRB | 2296 | 2305 | + | 18.10 | TCAAGGTCAT |
| L1MC4a_3end | ZNF140 | 365 | 383 | - | 17.46 | AGAAGTGGGATTGCTGGGT |
| L1MC4a_3end | DOF5.1 | 625 | 643 | + | 17.39 | AAAAAGCAAAGTGCAAAAG |
| L1MC4a_3end | THRB | 356 | 368 | - | 17.38 | GGGTCAAAGGGTA |
| L1MC4a_3end | DOF1.5 | 2611 | 2626 | + | 17.09 | AAAATAAAAAGTTAAA |
| L1MC4a_3end | CG7928 | 1564 | 1575 | - | 16.92 | CCCAGCCCTGGA |
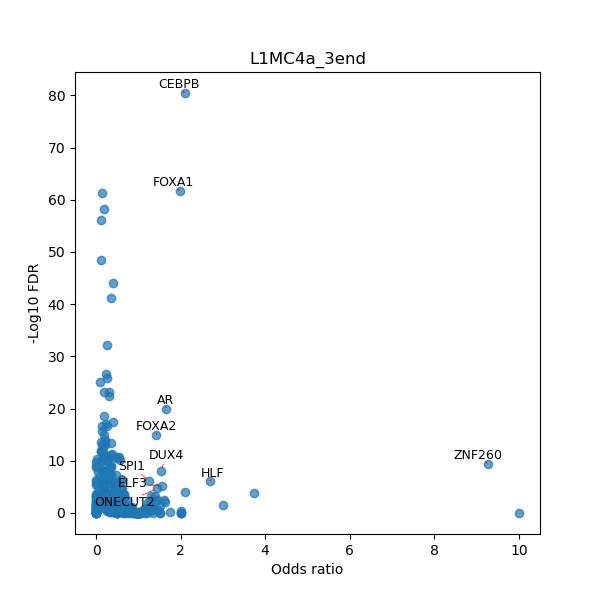
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.