L1M3e_5end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000242 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Euarchontoglires |
| Length | 2113 |
| Kimura value | 24.19 |
| Tau index | 0.9892 |
| Description | 5' end of L1 retrotransposon, L1M3e_5end subfamily |
| Comment | Contains multiple ~30 bp tandem duplications in the promoter region. |
| Sequence |
GACGTCAGCAAGATGGCGGAATAGGACTTTCCAGCGCTCGTCCTCACAGAAACATCAATTTGAACAACTATCCACGCACGAAAATACCTTCACAAGAGCTAAGGAAACCAGGTGAGAGATTACAGCACCTGGGTGTAGCACAGAAATAAGAAAAGACGCATTGAAGAGGGTAGGAAGGACAGTTTTACATTACCCGCGTCACCCCTCCCCCAACCCCAGGCAGCACAGCGTGGAGAGAGATACCCTCCGCNTGGGGGAAGGAGAGGGAAGTGAGCACCGGACTTTGCCTCGGACCCCAANCACCGGGCCCGCCCCAGTAAAACCCAGCGCCGGGCAGGCCCCCACGGCCCCAGACTCCAGGCCGGTACCCGCGGACTGAGCCGCGCCAGATCCCACAGCCCAGGCTCCAGGCCTGCCCGGCGGACTCAGTCTCCAGGCCCGCCCCAGCGCCAGGCCGACCCCAGTGCCCCAGGCTCCAGACCGGCCCCAGCGCCAGGCCGGCCCCAGCGGCCCCAGGCTCCAGGCCGNCCCCAGCGCCAGGCCGGCCCCCGCGGCCCCAGGCTTCAGGCCCGCCCCAGCACCAGGCCGGCCCCCGCAGCCCTAGNCATCAGGCCAGCACCCGCGGACCCAGCCTCCAGGCCGGCCCCTGCGGACACAGGCTCCAGGCCTACCCAGCGCCAGGCCAGCCCCTGCGGCCCCAGGCTCCAGGCCCACCCCAGGTTCCAGACCAGCCCAGAGCCAGGTCGGCCCACGTAGCCCCAGGCTTCAGGCCCGCCCCAGCGCCAGGTCAGCACCCCTGGCCTCAGACCNCGNCCCGGTACCAGGCCGGCACCCGCGGACACAGGCTCCAGGCCTGCCCAGTGCCAGGCCGGTCCCTGCGGCCCCACCCTCCAGGNCCGGCCCCTGCGGCCCCACGCTCCAGCAGACCCAGGGTCCAGGCCCGTCCCAGTAGACCCCAGCGCTAGGCCGGCCCCCACGGACCCAGGCTCCAGGACCGCCCCTGTGGACCCAGGTCCCAGGCCGGCCCCCGCGGCCCCAGGACCCAGGCCAGCCCTCGNAGACCTAGCCTCCAGGCCAGCCCTGCAGACCCAGCCTCCAGGCCGGCNCCCGCGGACCCAGGCTCCAGGCCATCCCCCAGGCTCCAGGCCAGCCCCAGTGGCTCCAGGCACCAGGCCAGCNCCCGCGGACCCAGGCTCCAGACCGGCCCCACGCTACCCCAGCACCAGGCCAGCCCCAGGCTCCAGGCCGGTCCCCGTGGCCCAGGCTCCAGTGGACCCAGGGTCCAGGCCTGCTCCAGCAGACCCAGGGTCCAGGCCCACCCCAGTAGACCCCGGCGCCAGGCCGGCCCCCGCGGACCCAGGCTCCAGGACCACCCCTGCAGACCCAGGCTCCAGGCCAGCCCCCGCGGACCAGGATCCANGGCCCANNNCCNCAGNCGCAGGCTCCAGGCCCGCCCCAGCGCCAGGCCAGCCCCCATGGACTCAGGCTCCAGGCCCGTCCCAGTGGACCCAGGCNCCAGGCCCATCCCAGCGCCAGGCCGGCCCCTGCAGACTCAGGCTCAAGGCCCACCCCAGCGCCAGGTCAGCCCCTGTGGACCCAGGCTTCAGGCCGGCCCCCGCGGACNCAGGCTCCAGGCCCGCCCTCGTGGACCCAGGCTCCAGGCCCACCCCCGCGGACCCAATCAACAGGTCCACCCAGTGGATCCAGGCTCCAGGCCCAACCCCGCGGACCCAGGCGCCAGGCCCGCCACCTGCTGACCCAGGCACCAGGCCAGCCTGCCCGAGGACTCCAGCAGCAAGCCCGCCCGCGGACCACGCCAGACGGCCTGCCCAGAATCTCTGGATGACTGGTGAAGGGCTTTCCCNGACAAAGCCAGTCTGCAAAGACTGGAATAAGTNCCTACTTCTTCAAATGCGCAGACACCAACGCAAGGCCCAAGGATCACGAANAATCAGGGAAACATGACACCACCAAAGGAACAAAATAAAGCGCCAGTAACCGACCCTAAAGAAATGGAGATNTACGAACTGCCTGACAAAGAATTCAAAATAATCGTTTTAAGGAAGCTCAGTGAGCTNCAAGAGAACACAGATAAACAACTAAACGAAATCAG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1M3e_5end | ESR1 | 1768 | 1782 | - | 0.33 | CGGGCAGGCTGGCCT |
| L1M3e_5end | ESR1 | 1954 | 1968 | + | 0.21 | AGGGAAACATGACAC |
| L1M3e_5end | RARA | 167 | 184 | + | -4.41 | AGGGTAGGAAGGACAGTT |
| L1M3e_5end | GLIS1 | 742 | 756 | + | -7.91 | GGTCGGCCCACGTAG |
| L1M3e_5end | EWSR1-FLI1 | 165 | 182 | + | -11.17 | AGAGGGTAGGAAGGACAG |
| L1M3e_5end | DAL81 | 1797 | 1815 | - | -38.59 | GTGGTCCGCGGGCGGGCTT |
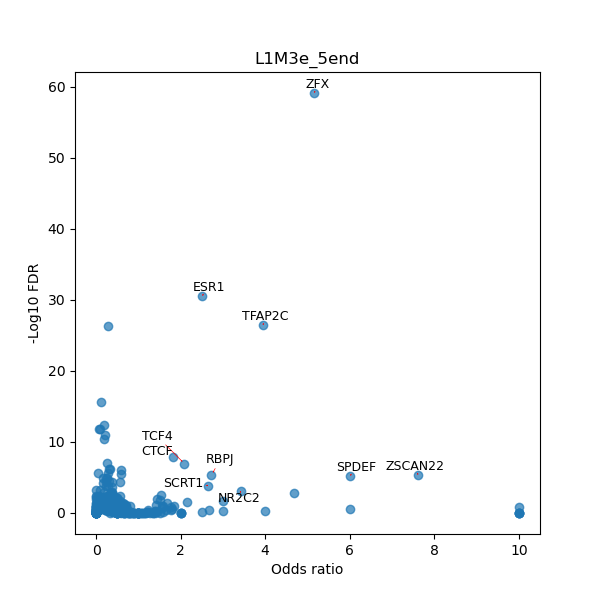
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.