L1M3e_5end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000242 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Euarchontoglires |
| Length | 2113 |
| Kimura value | 24.19 |
| Tau index | 0.9892 |
| Description | 5' end of L1 retrotransposon, L1M3e_5end subfamily |
| Comment | Contains multiple ~30 bp tandem duplications in the promoter region. |
| Sequence |
GACGTCAGCAAGATGGCGGAATAGGACTTTCCAGCGCTCGTCCTCACAGAAACATCAATTTGAACAACTATCCACGCACGAAAATACCTTCACAAGAGCTAAGGAAACCAGGTGAGAGATTACAGCACCTGGGTGTAGCACAGAAATAAGAAAAGACGCATTGAAGAGGGTAGGAAGGACAGTTTTACATTACCCGCGTCACCCCTCCCCCAACCCCAGGCAGCACAGCGTGGAGAGAGATACCCTCCGCNTGGGGGAAGGAGAGGGAAGTGAGCACCGGACTTTGCCTCGGACCCCAANCACCGGGCCCGCCCCAGTAAAACCCAGCGCCGGGCAGGCCCCCACGGCCCCAGACTCCAGGCCGGTACCCGCGGACTGAGCCGCGCCAGATCCCACAGCCCAGGCTCCAGGCCTGCCCGGCGGACTCAGTCTCCAGGCCCGCCCCAGCGCCAGGCCGACCCCAGTGCCCCAGGCTCCAGACCGGCCCCAGCGCCAGGCCGGCCCCAGCGGCCCCAGGCTCCAGGCCGNCCCCAGCGCCAGGCCGGCCCCCGCGGCCCCAGGCTTCAGGCCCGCCCCAGCACCAGGCCGGCCCCCGCAGCCCTAGNCATCAGGCCAGCACCCGCGGACCCAGCCTCCAGGCCGGCCCCTGCGGACACAGGCTCCAGGCCTACCCAGCGCCAGGCCAGCCCCTGCGGCCCCAGGCTCCAGGCCCACCCCAGGTTCCAGACCAGCCCAGAGCCAGGTCGGCCCACGTAGCCCCAGGCTTCAGGCCCGCCCCAGCGCCAGGTCAGCACCCCTGGCCTCAGACCNCGNCCCGGTACCAGGCCGGCACCCGCGGACACAGGCTCCAGGCCTGCCCAGTGCCAGGCCGGTCCCTGCGGCCCCACCCTCCAGGNCCGGCCCCTGCGGCCCCACGCTCCAGCAGACCCAGGGTCCAGGCCCGTCCCAGTAGACCCCAGCGCTAGGCCGGCCCCCACGGACCCAGGCTCCAGGACCGCCCCTGTGGACCCAGGTCCCAGGCCGGCCCCCGCGGCCCCAGGACCCAGGCCAGCCCTCGNAGACCTAGCCTCCAGGCCAGCCCTGCAGACCCAGCCTCCAGGCCGGCNCCCGCGGACCCAGGCTCCAGGCCATCCCCCAGGCTCCAGGCCAGCCCCAGTGGCTCCAGGCACCAGGCCAGCNCCCGCGGACCCAGGCTCCAGACCGGCCCCACGCTACCCCAGCACCAGGCCAGCCCCAGGCTCCAGGCCGGTCCCCGTGGCCCAGGCTCCAGTGGACCCAGGGTCCAGGCCTGCTCCAGCAGACCCAGGGTCCAGGCCCACCCCAGTAGACCCCGGCGCCAGGCCGGCCCCCGCGGACCCAGGCTCCAGGACCACCCCTGCAGACCCAGGCTCCAGGCCAGCCCCCGCGGACCAGGATCCANGGCCCANNNCCNCAGNCGCAGGCTCCAGGCCCGCCCCAGCGCCAGGCCAGCCCCCATGGACTCAGGCTCCAGGCCCGTCCCAGTGGACCCAGGCNCCAGGCCCATCCCAGCGCCAGGCCGGCCCCTGCAGACTCAGGCTCAAGGCCCACCCCAGCGCCAGGTCAGCCCCTGTGGACCCAGGCTTCAGGCCGGCCCCCGCGGACNCAGGCTCCAGGCCCGCCCTCGTGGACCCAGGCTCCAGGCCCACCCCCGCGGACCCAATCAACAGGTCCACCCAGTGGATCCAGGCTCCAGGCCCAACCCCGCGGACCCAGGCGCCAGGCCCGCCACCTGCTGACCCAGGCACCAGGCCAGCCTGCCCGAGGACTCCAGCAGCAAGCCCGCCCGCGGACCACGCCAGACGGCCTGCCCAGAATCTCTGGATGACTGGTGAAGGGCTTTCCCNGACAAAGCCAGTCTGCAAAGACTGGAATAAGTNCCTACTTCTTCAAATGCGCAGACACCAACGCAAGGCCCAAGGATCACGAANAATCAGGGAAACATGACACCACCAAAGGAACAAAATAAAGCGCCAGTAACCGACCCTAAAGAAATGGAGATNTACGAACTGCCTGACAAAGAATTCAAAATAATCGTTTTAAGGAAGCTCAGTGAGCTNCAAGAGAACACAGATAAACAACTAAACGAAATCAG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1M3e_5end | ZNF281 | 202 | 211 | - | 20.07 | GGGGGAGGGG |
| L1M3e_5end | ZNF85 | 116 | 127 | + | 19.71 | GAGATTACAGCA |
| L1M3e_5end | ZNF148 | 202 | 211 | + | 18.97 | CCCCTCCCCC |
| L1M3e_5end | phol | 9 | 18 | - | 18.44 | CGCCATCTTG |
| L1M3e_5end | ZKSCAN3 | 1042 | 1055 | + | 18.37 | CCCAGGCCAGCCCT |
| L1M3e_5end | Klf15 | 193 | 214 | + | 18.30 | CCCGCGTCACCCCTCCCCCAAC |
| L1M3e_5end | ZBTB24 | 1035 | 1044 | + | 17.49 | CCCAGGACCC |
| L1M3e_5end | CTCF | 1744 | 1758 | - | 17.39 | GTCAGCAGGTGGCGG |
| L1M3e_5end | klu | 1664 | 1674 | + | 16.33 | CCACCCCCGCG |
| L1M3e_5end | ZBTB6 | 1556 | 1564 | - | 16.28 | CCTTGAGCC |
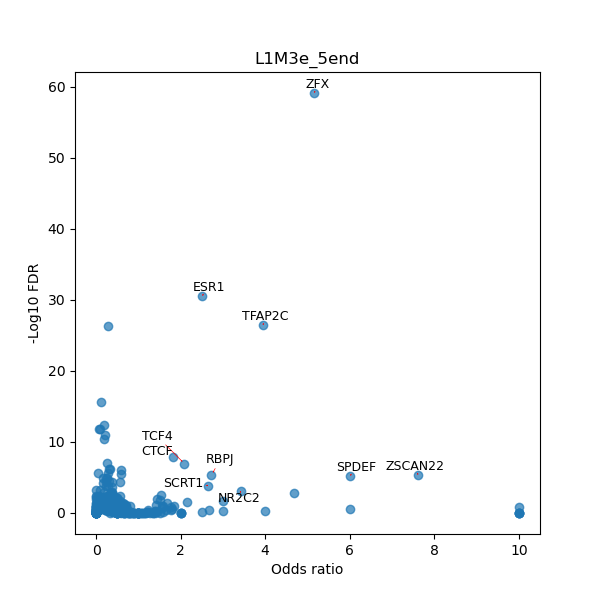
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.