HAL1ME
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000157 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Eutheria |
| Length | 2465 |
| Kimura value | 32.00 |
| Tau index | 0.8302 |
| Description | L1 retrotransposon, HAL1ME subfamily |
| Comment | Full ORF1 between ~724-1707. |
| Sequence |
GGCTTCCGGTTCAAGATGGCGGACTGAGCACACGCGTCTACCTCCCCTCCCTCCCAAAATCCCATTGAAATGACAGAAAAGATATTTTTAAAAGAATGAAACCATAACAGCGCTGGAAAACAAGAAAGGGTGCCATCAGCGGACCAGAAATTTTGAGGAATTCCTGGAAGATAGAAAGCAGATGGGATCGGATTGACGGAGAAANCAGAGCAGAGGAAACCGCAGCCCAAAACACCGGAGGTAAGAGCGGTTCTTCCCAAGGGAGCCCCGGAGAAGCTCCAAGCTCGGAGTCAACAAATACAAAGAGCAGGAGTGGGCCGTGGGGCAATCATCAGGGTAATTAATTAAAGAACTGCATTCGGAACAGCTGGGCCAGTGGGTCCTCCCTCCCCCTCTCCCGCGTANACANAGAGCAGCTGGCAGCAGCGGCTTTTGCCCCCAGGCTAAAGCCCGAGAACTGCTCTCTAAAGAAAGTGGGCGATCTGCCTGGGGAGGGCTCCTAGTGTGGGTGTTGGGGCCTGAGAGAGAAGCAGAGCAGAGCTTGAGGTAACTCCGGACCTCCACAACAGACACGGAGGGGAAAAAGGAAAGGGAAGAGGCAGCTCCGCCCAGGAGTGAAATACACTAAAGCGCTCCACCCCCAGAGTAAAGCCCTCCTTACTTTGGCAATTGTGGGGGTCCCCAGCCTGCCTGCCTGCTCCCTGCCCACGCACCCTAAAGGGAAGCCTGCCTGTCGACACGCCCCGCCCACTTACACAGGAGCCCGCAGTCAGCCCTCCCATTCAAGGGAGAGGCCTGTTGGGGAAACAGACCTACACAACTGCAGAGGAAACCCACGCCAGCTACCTATACTAATCAACCTAGTCCTTCATTCATAAATATGAACGGACAACCAAGGATCACCAGACATTTGAGGAAAACCAACAGCATGAAAGAGAAGGACCAAGATGAACAAACAGAACAACTGACCCCGGAGGAAACAGAGATAATTCAGGGAACAGAAGAGAACTTTAAAAAAATTCTAATTAGTATCCTCAGAGAGATTCGAGAAGATATTGCATCCATAAAACAAGAACAGGNTGCTATGAAAAAGAAGCAATCAGAGAACAAGAAAGAGTTCTTGGAAATTAAAAATATGATTGCCAAAATNAAAAATTCAATAGAAGGGTTGGAAGATAAAGTCGAGGAAATCTCCCAGAACGTAGAGCAAAAAGACAAAGAGATGGAAAATATGAGAGAAAAGTTAAGAGACATGGAGGATAGATCCAGGAGGTCCAACATCCATCTAATAGGAGTTCCAGAAGGAGAGAACAGAGANAATGGAGGGGAGGAAATAATCAAAGAAATAATAGAAGAAAATTTCCCNGAGCTGAAGAAAGACACGAGTCTTCAGATTGAAAGGGCCCACCGAGTGCCGAGCAGGATGAATGAAAAAAGACCCACACCTAGACACATCNTNGTGAAATTTCAGAACNCCAAGGATAAAGAGAAAATCCTAAAAGCTTCCAGAGAGAAAAAACAGGTTACCTACAAAGGAACGAGAATCAGACTGGCATCAGACTTCTCATCAGCAACACTGGATGCTAGAAGACAATGGAGCAATGTCTTCAAAGTTCTGAGGGAAAATNATTTTGAACCTAGAATTCTATACCCAGCCAAACTATCAATCAAGTGTGAGGGCAAAATAAAGACATTTTCAGACATGCAAGGACTCAGAAAGTTTACCNCCCACGTACCCTNTNTGAAAAAATTACTCGAGGATGTACTCCAGCAAAACGAAAAAGGAATCCAAGAAAGAGGANGACATGGGATACAAGAAACAGTGGCAGCTGAGTATGACCCAGAAGAAGTGAAGGAAGCTAAAAAGAAATTAAAAGAGAATCCATTTGACCTTGACGCTTGGAANATTCTCCTTCGAGCGGCANAGGTTTAGTGACATAGGATTACATTTCCTTCTCTATGGGTCCAATCACACTACTTGGTTCTGCAGTGAATAATATTTACATAATCATAATANTGTAAATGCTGTTTATTGGTTTTCAATTTTTAGAATCAACCTATAGACAAAGCACGGAAGACTTAATTATAGTTACAGAACAGAATGTAAATGTTATAAACCTTGACAATGTAAAAATAATANAATAACNAAAATTGGGAGGTAGAAGGGGGAAGAGAGGTGGAAGGAAGTGTAAGNGCGCTAANTTCCTCATCTTNCATAGCGGGGAGTCAANAGATACTGTCTAAAGTTGATAAATCAAGAAATAGAGGTNTAAGTATATTATTTAAAGTTATAAAGGTAACCANTAGAAGAACTAAAAATAANTAACAGTNAAAAGTGGTTGCCTCTGGGGAGTGGGATNGAGATGGAAGAGGGNAGGGANAGGGACTACTTTTCATTTTATACCCTTCTGTACTGTTTGAATTTTTTNNACCATGTGCATGTATTACTTTTATAAAAATAAAAA
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| HAL1ME | KLF14 | 742 | 750 | - | 16.15 | TGGGCGGGG |
| HAL1ME | Nr5A2 | 1888 | 1896 | + | 16.13 | TGACCTTGA |
| HAL1ME | DOF3.6 | 1227 | 1247 | - | 16.11 | TTAACTTTTCTCTCATATTTT |
| HAL1ME | MAZ | 44 | 51 | + | 16.08 | CCCCTCCC |
| HAL1ME | ZNF148 | 44 | 53 | + | 16.08 | CCCCTCCCTC |
| HAL1ME | RVE8 | 80 | 88 | + | 16.08 | AGATATTTT |
| HAL1ME | TFAP2A | 434 | 444 | + | 16.05 | TGCCCCCAGGC |
| HAL1ME | LjSGA_053525.1 | 1401 | 1408 | + | 16.05 | GGGCCCAC |
| HAL1ME | sta-1 | 1118 | 1126 | - | 16.04 | TTCCAAGAA |
| HAL1ME | ETS2 | 2 | 10 | - | 16.00 | ACCGGAAGC |
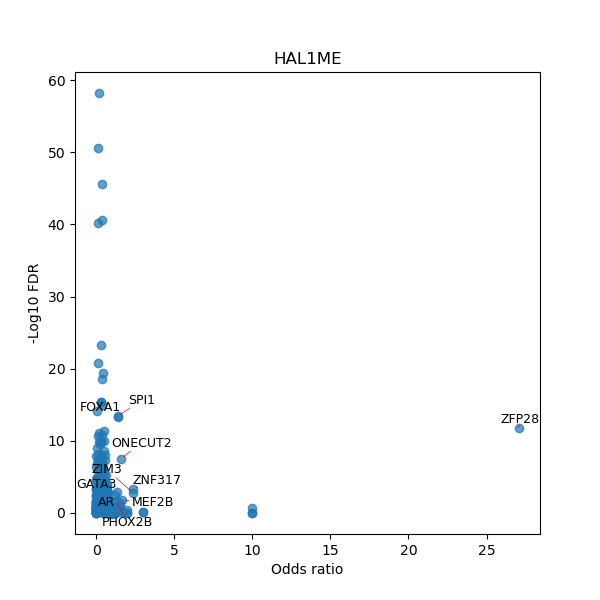
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.