Eutr2B
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001264 |
|---|---|
| TE superfamily | MuDR |
| TE class | DNA |
| Species | Eutheria |
| Length | 1699 |
| Kimura value | 25.24 |
| Tau index | 0.0000 |
| Description | Eutr2B subfamily |
| Comment | Pos 13-125 bp of the 250-300 bp long TIRs are ~8 copies of ATTCGCCGAAARYCA. Full TIRs are shared with the shorter element named Eutr2. The remaining 1200 bp are unrelated. |
| Sequence |
GGCTCTAGGCCAATTCGCCGAAAATCAATTCGCCGAAAGCCAATTCGCCGAAAGATCAATTCGCCGAATGACCAATTCGCCGAAAGCCAATTCGCCGAAAATCAATTTGCCGAATGACCAATTCGCCGAATTTACCAAATTTACCGATTTACCAAAAACTTGTTTCTTTTAACTATTTATAAAGTTTACAGCAATTCGTATTGGATGGGATGGTTTTAACAGCTTTTGAAGATTTCTGCGAAGTTTTAGTTGACCGGTTTTCATTTTGTCTTCATAGCAGCATTTTAAGGAATGTTCAATTTGTCGAAAGCTAAGGGTCATAGGGTGGTTTAGTCTATTTNTGTTAAATATATTTAATATTCAGAAAAGTATATTTTATCATCAGTAACCAAAATTCAGCATTTCATATGGGAATTTTTCAGTTTATACTGATTTCTTTCNAATNTTTTTGAAACTTTTTATNATCCATTTTTATGCTTTTTTNAATTATTTTNAGGATTCTTTCGCCAATTTCTCAAGTAGCATATCAACCTTAGTTTGTTATTTCTAGNAATTAGTAGCCAAAATTTAATATGTGATGTGGTTGCTCCTGTCGCTTCCTAGTTCATTTTACTTACCTATCTTTTAGCATTTTTAGGATTTTTAANANTTAATTTTTGATCCTTCCTTCACAATTTCGTATTTATGGTTAATGTATATCAAATTTGAGGTACTAATCATACGGTCCATTATAGTAATCCCTTCTGGGCATCCTCACTTCCTTTCCGCTGCTTTGAACGGTTTTACCCTTTCATGTCGAATCTTTCTGCAAACTGCTACCAAGTCGACATATCCCAATTGCTCCTTGTTAGAAACCAAAATTTCAGCATTTCATATGTGACGGGATTGTTTCGGCTACTTCTGATTTCTTTCGGATGCTTTTGAATGTCATTTTCTATTTTTTCAAACTTTAANCATTTTTAGTATTCTTTCGCCAATTCCTCAAATGGCATATCAGCCTTAATTTTGTATTTCTAGCAATTAAAATTTTAAATTTCAGCATTTCACATGTGATGGGATTGTTTCGGCCGCTTTTGAGTTCTTTCGCATCCTTTTGAATGTCATTTTTNATTTTTCCAATCTTTAGGCATTTTTACCATTCTTTCACCAATGTCTCAAATGGCATATCAGCCTTAATTTTGTATTTCTAGCAATTAGTAACCAAAATTTCGGCATTTCATATTGATGTGGTTGCTCTGGATCCTTCCTCAGCAATTTCATATTTATGGTGAATGTATATCAGATTTGGGGTACTAATCTTACGCTGAAATTTTGGTTACTGATTATGAAATATACTTTTCTGAATATTAAATATATTCATAAGAAGACTAAACCACCCTATGACCCTTAGCTTTTGACAAATTGAACATTCCTTAAAATGCTGCTATGAAGACAAAATGAAAACCGGTCAACTAAAACTTCGCAGAAATCTTCAAAAGCTGTTAAAACCATCCCATCCAATACGAATTGCTGTAAACTTTATAAATAGTTAAAAGAAACAAGTTTTTGGTAAATCGGTAAATTTGGTAAATTCGGCGAATTGGTCATTCGGCGAATTGATTTTCGGCGAATTGGCTTTCGGCGAATTGGTCATTCGGCGAATTGATTTTCGGCGAATTGGCTTTCGGCGAATTGATTTTCGGCGAATTGGCCCGGAGCC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Eutr2B | DOF5.1 | 160 | 178 | - | 14.30 | AAATAGTTAAAAGAAACAA |
| Eutr2B | Dlx5 | 1190 | 1197 | + | 14.30 | GCAATTAG |
| Eutr2B | Elf5 | 756 | 763 | - | 14.24 | AAGGAAGT |
| Eutr2B | vnd | 513 | 521 | + | 14.22 | TCTCAAGTA |
| Eutr2B | MYB80 | 131 | 142 | + | 14.11 | TTTACCAAATTT |
| Eutr2B | MYB80 | 1559 | 1570 | - | 14.11 | TTTACCAAATTT |
| Eutr2B | MAFF | 1247 | 1257 | + | 14.04 | CTCAGCAATTT |
| Eutr2B | MGP | 1426 | 1438 | + | 14.03 | ATGAAGACAAAAT |
| Eutr2B | MGP | 263 | 275 | - | 14.03 | ATGAAGACAAAAT |
| Eutr2B | THRB | 1374 | 1386 | - | 13.99 | GGGTCATAGGGTG |
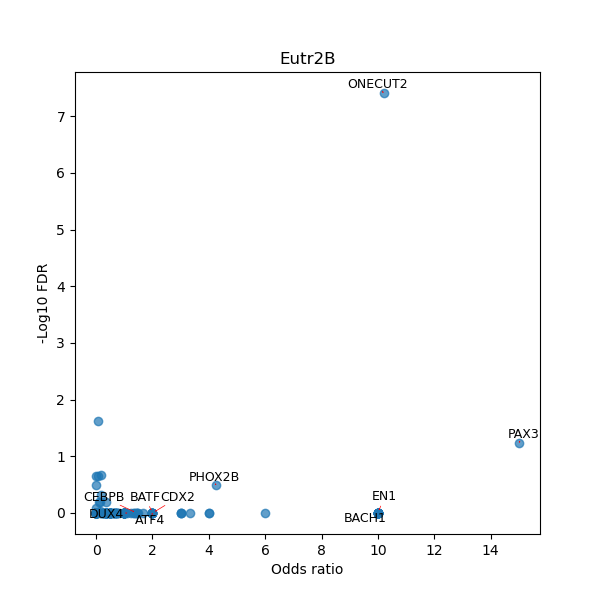
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.