Eutr2B
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001264 |
|---|---|
| TE superfamily | MuDR |
| TE class | DNA |
| Species | Eutheria |
| Length | 1699 |
| Kimura value | 25.24 |
| Tau index | 0.0000 |
| Description | Eutr2B subfamily |
| Comment | Pos 13-125 bp of the 250-300 bp long TIRs are ~8 copies of ATTCGCCGAAARYCA. Full TIRs are shared with the shorter element named Eutr2. The remaining 1200 bp are unrelated. |
| Sequence |
GGCTCTAGGCCAATTCGCCGAAAATCAATTCGCCGAAAGCCAATTCGCCGAAAGATCAATTCGCCGAATGACCAATTCGCCGAAAGCCAATTCGCCGAAAATCAATTTGCCGAATGACCAATTCGCCGAATTTACCAAATTTACCGATTTACCAAAAACTTGTTTCTTTTAACTATTTATAAAGTTTACAGCAATTCGTATTGGATGGGATGGTTTTAACAGCTTTTGAAGATTTCTGCGAAGTTTTAGTTGACCGGTTTTCATTTTGTCTTCATAGCAGCATTTTAAGGAATGTTCAATTTGTCGAAAGCTAAGGGTCATAGGGTGGTTTAGTCTATTTNTGTTAAATATATTTAATATTCAGAAAAGTATATTTTATCATCAGTAACCAAAATTCAGCATTTCATATGGGAATTTTTCAGTTTATACTGATTTCTTTCNAATNTTTTTGAAACTTTTTATNATCCATTTTTATGCTTTTTTNAATTATTTTNAGGATTCTTTCGCCAATTTCTCAAGTAGCATATCAACCTTAGTTTGTTATTTCTAGNAATTAGTAGCCAAAATTTAATATGTGATGTGGTTGCTCCTGTCGCTTCCTAGTTCATTTTACTTACCTATCTTTTAGCATTTTTAGGATTTTTAANANTTAATTTTTGATCCTTCCTTCACAATTTCGTATTTATGGTTAATGTATATCAAATTTGAGGTACTAATCATACGGTCCATTATAGTAATCCCTTCTGGGCATCCTCACTTCCTTTCCGCTGCTTTGAACGGTTTTACCCTTTCATGTCGAATCTTTCTGCAAACTGCTACCAAGTCGACATATCCCAATTGCTCCTTGTTAGAAACCAAAATTTCAGCATTTCATATGTGACGGGATTGTTTCGGCTACTTCTGATTTCTTTCGGATGCTTTTGAATGTCATTTTCTATTTTTTCAAACTTTAANCATTTTTAGTATTCTTTCGCCAATTCCTCAAATGGCATATCAGCCTTAATTTTGTATTTCTAGCAATTAAAATTTTAAATTTCAGCATTTCACATGTGATGGGATTGTTTCGGCCGCTTTTGAGTTCTTTCGCATCCTTTTGAATGTCATTTTTNATTTTTCCAATCTTTAGGCATTTTTACCATTCTTTCACCAATGTCTCAAATGGCATATCAGCCTTAATTTTGTATTTCTAGCAATTAGTAACCAAAATTTCGGCATTTCATATTGATGTGGTTGCTCTGGATCCTTCCTCAGCAATTTCATATTTATGGTGAATGTATATCAGATTTGGGGTACTAATCTTACGCTGAAATTTTGGTTACTGATTATGAAATATACTTTTCTGAATATTAAATATATTCATAAGAAGACTAAACCACCCTATGACCCTTAGCTTTTGACAAATTGAACATTCCTTAAAATGCTGCTATGAAGACAAAATGAAAACCGGTCAACTAAAACTTCGCAGAAATCTTCAAAAGCTGTTAAAACCATCCCATCCAATACGAATTGCTGTAAACTTTATAAATAGTTAAAAGAAACAAGTTTTTGGTAAATCGGTAAATTTGGTAAATTCGGCGAATTGGTCATTCGGCGAATTGATTTTCGGCGAATTGGCTTTCGGCGAATTGGTCATTCGGCGAATTGATTTTCGGCGAATTGGCTTTCGGCGAATTGATTTTCGGCGAATTGGCCCGGAGCC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Eutr2B | TCX6 | 447 | 461 | + | 15.15 | TTTTGAAACTTTTTA |
| Eutr2B | STB3 | 413 | 423 | + | 14.92 | AATTTTTCAGT |
| Eutr2B | JKD | 1426 | 1437 | - | 14.76 | TTTTGTCTTCAT |
| Eutr2B | JKD | 264 | 275 | + | 14.76 | TTTTGTCTTCAT |
| Eutr2B | IDD1 | 1426 | 1437 | - | 14.56 | TTTTGTCTTCAT |
| Eutr2B | IDD1 | 264 | 275 | + | 14.56 | TTTTGTCTTCAT |
| Eutr2B | NR2F1 | 1377 | 1388 | - | 14.40 | AAGGGTCATAGG |
| Eutr2B | NR2F1 | 313 | 324 | + | 14.40 | AAGGGTCATAGG |
| Eutr2B | Dlx2 | 1190 | 1197 | + | 14.32 | GCAATTAG |
| Eutr2B | DOF5.1 | 1523 | 1541 | + | 14.30 | AAATAGTTAAAAGAAACAA |
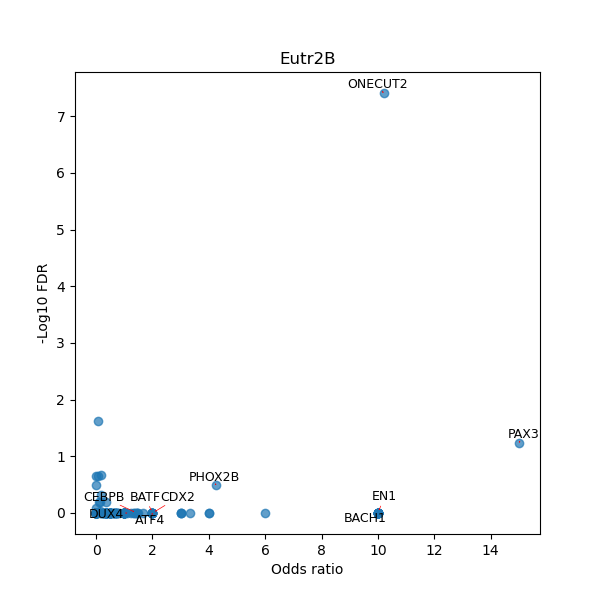
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.