ERV3-16A3_I
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000116 |
|---|---|
| TE superfamily | ERV3 |
| TE class | LTR |
| Species | Theria_mammals |
| Length | 5226 |
| Kimura value | 31.08 |
| Tau index | 0.8928 |
| Description | Internal region of an ERVL endogenous retrovirus, ERV3-16A3_I subfamily |
| Comment | Reconstructed from the human genome, but it's shared by all placental mammals. |
| Sequence |
CATGAGTGGTACCGGGAGTGGTCCGNAGAAAACAGGCGGTAAGATGGGGTTTTGGGACTGGCTCACTCACCGCCCGGCGGACAGAGGATACCATCCTGGCACAAGGTGGCGGCCCAGTTGCTAAAGATTTCACCGGTGGTGACCTGGGAAAATGTGTCCCAGNNNTTGGAAGGGGAATGCCCTTGCAGGTGCAATGATTCAGGCATTTGAAAGATATGGGGAGANAATGCCTACAAGGACAGCGGAGTTGGCTGGTTATTGCTAAGTTGTATTGATGCCCTGCAGAGGGATAATGAGAAACTGAGGGCNGTTAACAAGCAGTTAAAGGCTAAGTGTGAGAGCCAGAGGGCCTCTTTGNTGGTAGCTTACAAAGAGGCCCTTATCTCCTGCAGTGGAAGAGCAGACANAGCTGAGGAGCAGACTCAGGATCTAATAGTTAGAGTCGCAGAGCTCCAGAGACGTTTGAATGCTCAGCCAAGGCGAGGTCTGTTATGCTAAAGNTAAGGCCCTTGGTTGGGAAAACCTGGGACGGAACACGGGATGGGGACATCTGGGTGGATGCCCCCGAAGATNTTGGCTCTCCAGACTCCTCTGAACCCTCNGAGCCTGCAGAAGTGGCCCACCCCTCCCTANTAAGAGCTAGCACTCCTCTTGCGCCCGCGCTGGAAGACGNTGCAGAGGCCTCTCCCCGCAAGGCAACAGGTGCCCCCTCTCAGGANCTGCCCCCACCTCCCCTCCTGGCCGCCAGGCCGATAACTAGGGTTAAGTCNCAGCATAACCCGGCTGGGGACGTGCTGGGCCTGATAAGGGAGGAAAGGGACTATACCCCAAAGGAGCTGCAAGAATTAGCCAGCATGTACCGGCAGGAGCCAGGGGAGTATACNCNTGGGACTGGATTTTGAGGGTGCTTGATCAAGGGGGCCGGAATATAAGCGGTAAGACTGGATAANGGAAGAATTTATTGACTTGGGAGCACTTTCTCGGGATACAGGATTTAACACCCTGGCAAGGACCCCAGGAGATGGTGCAAACTCGCTGCTAGGATGGCTCCTAGAAGCGAGAAAAATGGCCCACGCTGAGCGAAGTNGAAATGCCNGAATTGCCNTGGCAGACGGTAGAGGAAGGGATTAAAGGCTCAGGGAAGTGGGCATGCTGGAATGGATATNATTATGTAAGGCCGGAAGACCCACCAGANGATTATGTTCCACGGGAGGGCCCACACCATTCACCAAGGCCATNAGGAATGCGCTGGTGAGAGGGGCACCAGCANNTCACTAAGAAGTTCAGTGGTGGCTCTCCTCTGCAGGCCAGGGCTGACGGTAGGAGAGGCCGTTACAGAACTGGGCTCGCTGATAGCAATGGGGATGATAGGACCCCGAAACAATAGAGGCCAGGTGGCGGCGCTTAACCGCCAGAAGCCAGGNGGNCGCAATTATCGTAATGACCGGAAGGTCGGAGTGGCAGCCAAGGGGGCCTGACCCAAGAGAGTTGTGGAGATGGTTAATAGAACACGGCGTCCCTAGGGGCAAAATAGATNGACAAGGGTGCTGCTTAACTTGTAACAGCAGAAAAAATCTGGGAGGCTGAGGGCGGTCGCCCCAATAAAAAAGTCACGATCCCTTGCCCAGTTTCCGGACCTGAGCCAGTTTTCAGACCCGGAACCCATTGACTGAAGAGGNGGCCGGGTCCCCAGGAGGAAGGACACTGCAATGATTCTTCTCNTTCCTCCCGNAGGGACCTATGGCCATTTACTCGGGTAACTGTACACTGGGGAAAGGGAATACCCAGACATTTCGAGGACTGTTGGACACAGGGTCTGAGTTGACATTGATACCCGGAGACCCGAAGCGTCATCATGGCCCCCTGTTAGAGTGGGGGCATATGGGGGCCAGGTAATAAATGGAGTCCTGGCCAAGGTCCGGCTCACAGTGGGTCCACTGGGTCCANGAACCCAGTGGTCATTTCCCCGGTCCCCGAATGTATAATTGGGTAGTTGGAGTAACCCCCACATTGGNTCCTTGGCCTGTGGGGTAAGAGCTATCATAGTGGGGAAGGCCAAGTGGAAGCCTCTGAAACTGCCCNCCNCCNCCATTCCTAGCCAAGATAGTAAATTATTATATCCTCTCGAGGAAGTGGCGGATGCAGATTAGTGCCACCNTTAAAGACCTAAAGGATGCAGGGGTGGTGGTCCCCATCATATCTCCATTTAATTCACCAGTCTGGCCCCTGCAGAAACCGGATGGATCCTGGAGAATGACNGTAGACTACCGCAAACTCAACCAAGTAGTAGCCCCGATTGCAGCTGCTGTGCCAGATGTGGTATCTTTGCTAGAGCAGATTAACACGGCCTCAGGTACATGGTATGCGGCCATTGATTTGGCGAATGCGTTCTTTTCCATTCCNATCAGAAAAGAGGATCAGAAACATTCACATGAACAGGAAANAATATACATTTACAGTTTTGCCCCAGGGCTATGTTATCTTCTCCTCTGTCATAATAAGTCCGAAGAGATCTGGACCGTCTGGACATCCCGCAGAACATCACATTGATCCATTACATCGATGACATCATGCTGATCGGACNGGATGAGCAAGAGGTGGCTACGTTAGGCCTTGGTAAGACACATGCGCTGAGGGTGGAGATAACCCTACGAAGATTCAGCGNNNGGACCTGCCACNTCAGTAAAGTTTTTAGGGGTCCAGTGGTCNGGGGCATGCCGGGACATCCCCTCCAAAGTAAAAGACAAATTGTTGCATCTTGCATCTCCTACCACGAAGAAGGAAGCACAACGCCTGGTAGGCCTCTTTGGGTTCTGGAGGCAGCATATTCCACACCTGGGAATACTGCTCCGGCCCATNTACCGGGTGACACGGAAGGCTGCCAGCTTTGAGTGGGCCCAGAGCAGGAAAGGGCTCTGCAGCAGGTCCAGGCTGCGGTGCAAGCAGCCCTGCCGCTTGGGCCATACGACCCGGCAGACCCTATGGTGTTGGAGGTGTCAGTGGTGGGAAAAGATGCNGTGTGGAGTTTATGGCAAGCCCCAGTGGGAGAATCACAACGCAGGCCCCTGGGGTTCTGGAGCAAGGCTGCAAGTCTTTTGAAAAACAGCTCTTNNNATGCTACTGGTAGAGANGGAACGCTTACCATGGGACACCAAGTGACACGNANCAAAATGTACCCCGTCATGAGCTGGGTTCTGTCGGACCCACCAAGTCATAAGGTCGGGCGGGCCCAGCAGCAATCCATCGTAAGATGGAAGTGGTACATCCGGGATCGAGCACGAGGAGGACCACAAGTAAGCTGCATGAGCAGGTAGCCCAGACCCCCATGTCACCCACCACNGTTGCACCAGCGCTCCTTCAGCTCACACCTATGGCCGTATGGAGNAGAGGTGTGNTCCTGATCGACCAGCTGAAGGAGGAGGAAAAANGCCCGAGCTTGGTTCACGGATGGGTCGGCTCGGTATGTGGGTGCAAGCCGAAAATGGACGGCGGCTTGCACTACAGCCNCACTCAGGGGTGGCCTTGAAAGACAGTGGNGAGGGAAAATCTTCCCAATGGGCAGAGCTTCGGGCGGTGCACCTGGTCATCCACTTTGTGTGGAAGGAGAAGTGGCCCGAGGTNAGAATATATACGGACTCATGGGCAGTGGCGAATGGCTTGGCCGGCTGGTCAGGGGCCTGGAAGGAGAAAGATTGGAAGATCGGAGACAAGGAGGTCTGGGGANTAGAGGCATGTGGATGGACNTATGGGAGTGGGCACGAAGTGTGAAGATCTTTGTATCACATGTTAACGCCCACCAGAGAGCATCCACCACGGAAGAGGCACTAAACAACCAAGTAGACAAAATGACTCGGCCAGTTGACGTCAGCCAGCCTCTGTCATCGGCCACCCCANGTGCTGGCACAATGGGCACATGAACGGAGTGGCCACGGTGGCAGAGATGGAGGCTACGCATGGGCCCAACAGCATGGACTCCCACTCACCAAGGCTGATCTAGCTACTTGCTGCCNCNNCTGAATGTCCAACCTGCCAGCAACAGAGACCGATCCCCGATATGGCACCATTCCTCGAGGAGACCAACCAGCCACTTGGTGGCAAGTTGACTACATTGGGCCCCTTCCATCCTGGAAGGGNCAGCGGTTCATCCTNACAGGAATACATATTCCGGGTATGGGTTTGCCTTTCCTGCCCGCAGGGCCTCAGCCAGCACCACTATCCGAGGGCTTACGGAGTGCCTGATCCACNGGCATGGGATCCCACACAACATNGCATCNGACCAGGGACCCACTTTACAGCAAAGGAGGTGCGGGAGTGGGCCCATGACCATGGGATCCACTGGTCNTATCACATACCGCACCATCCAGAAGCTGCCGGCCTGATAGAGCGNTGGAACGGCCTNCTGAAGGCACAGCTGAAGCGCCAGCTCGGAGACGATACTCTGCGAGGATGGGGCGCCATCCTTCAGGATGCAGTGTCCCCAATAGGAAGAATACATGGGTCCGGGAACCAAGGGGTGGAAGCAGGAGTGTNGCCCCACTTACCATCACTCCCAGTGACCCACTGGGGGANTTTGTGCTTCCCGTCCCCGCAACTCTGGGCTCTGCAGGNTTAGAGGTCCTGGTCCCCAAAGGGGGNACGCTCTCGCCAGGGGACACAGCGTCCCATTGAACTNTANAGCTACGGCTGCCGCCTGGGCACTTTGGGCTCCTTGTGTCCAGGGACCAGCAGGCAAGAAGAGGAGTCACCATCTTGGCAGGGGTAATTGACCCTGATCATCAGGAGGAGGTAGGGCTGCTNTTACACAATGGGGGCAGGGAGGAATACGTNTGGAACCCAGGTGATCCACTTGGGCGCCTCTTGGTACTCCCTTGCCCAATTNTGACTGTAAATGGACAAGTGCAGCAACCCCGGCCTGAGAAGGGCATGGTNACCAGGGGCTCAGACCCCTCAGGAATGAGGGTCTGGGTCACGCCACCAGGTAAGCCACCGAGACCAGCAGAGTGNTAGCTGAGGGTGAGGGGAATCTAGAATGGATAGTGGAGGAGGGAGACGATGAGTATCAGTTGCGGCCCCGAGACCAACTGAGCGACGGGGGCTGTAGTTCGTCCCACTAACCTCCCTCTTCTAAGTTTCCCTCAGGAAGAGAGGCCCACNGGAATCCTGGAGGAGCTGCTCCTGAACCGAACGTGTATGGAGAAGTGGATCCGNGCGGCGCAAGGGGTGGAC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| ERV3-16A3_I | JUN | 2563 | 2572 | - | 17.08 | ATGATGTCAT |
| ERV3-16A3_I | SCRT1 | 696 | 705 | + | 16.90 | GCAACAGGTG |
| ERV3-16A3_I | PRDM9 | 3922 | 3941 | + | 16.89 | AGTGGCCACGGTGGCAGAGA |
| ERV3-16A3_I | ZNF708 | 4404 | 4412 | - | 16.87 | GCTGTGCCT |
| ERV3-16A3_I | TGA6 | 2563 | 2572 | - | 16.80 | ATGATGTCAT |
| ERV3-16A3_I | IDD7 | 2736 | 2749 | + | 16.76 | AGTAAAAGACAAAT |
| ERV3-16A3_I | Ar | 1506 | 1521 | - | 16.74 | AGGGACGCCGTGTTCT |
| ERV3-16A3_I | CG12605 | 697 | 705 | - | 16.69 | CACCTGTTG |
| ERV3-16A3_I | TFAP2A | 4187 | 4199 | - | 16.59 | GGCCCTGCGGGCA |
| ERV3-16A3_I | ARALYDRAFT_496250 | 2185 | 2192 | - | 16.55 | GGGACCAC |
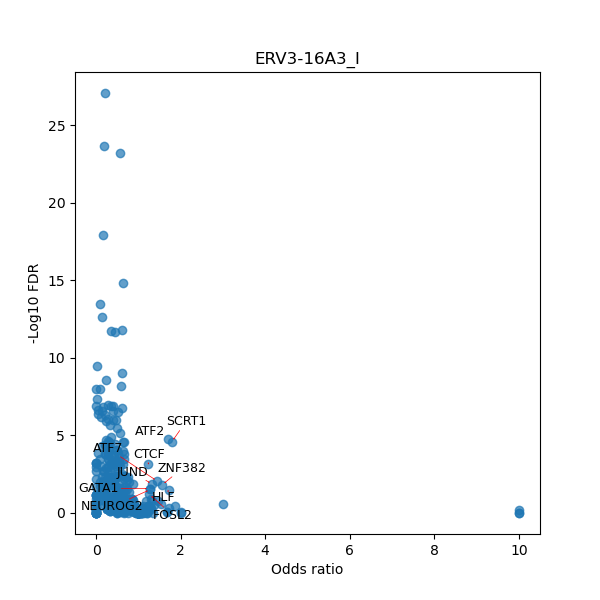
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.