Charlie8
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000107 |
|---|---|
| TE superfamily | Charlie |
| TE class | DNA |
| Species | Eutheria |
| Length | 2457 |
| Kimura value | 28.91 |
| Tau index | 1.0000 |
| Description | hAT-Charlie DNA transposon, Charlie8 subfamily (autonomous) |
| Comment | 15 bp TIR; 8bp TSD with MER1-like NTCTAGAN bias. Nearly complete transposase (roughly <330-2102). |
| Sequence |
CAGAGGTCGCAAACTGGCGGCCCGCGGGCCGNATCCGGCCCGCAGACGTGTTTTGTTTGGCCCGCACAGTGTTTTAAAANATTTTGAATTAGTTGCCAACATTTAAAAATCGGGAGATTTCACATAAAAATCCGGATTTCCGGCTTCTCTTGAAAAATCGGAAGATCTGGCAACACTGGGCCCGCATTCCCGCGTGGCAACAATCGGCTGGAGCTGAGTAGCGGCTGCCCCCTTTAGACGGGGCATGCGCTCTCCAGTTCGCCACAGTCCCCACCACTCCCTATTGTCTCCCCGACACTGAGGCCGAGTGTCAGTTGCCATTTATCATCGCGCTTGCGCTGTTGTTTTTCTTATAGTAGAGTTAAGAGGAAAGTGAAATATTTCTTGTACCCATGTCTCTATCAAAAGTGGGAAAACGAAAGATAGACCGAGAGGGCCGCGTGTTTCAAGAAAAATGGGAGAGAGCATATTTCTTTGTGGAAGTGAAGAATATTCCTACGTGTTTAATATGCAAACAAAGTGTGTCTGTGTCGAAAGAATACAACTTAAGACGCCATTATGAAACAAACCATAGTAAGAACTATGACCGGTATACGGAAAAGATGCGTGATGAAAAACTTAATGAACTGAAAAAAGGACTGAAATTTCAACAGGGTTTGTTTTTGAATGCGAATAAAATAAGTGATGCTGCTGTGAAATGCAGTTATGTATTAAGTGAAAAAATTGCCCGGGCATCAAAACCTTTTACAGATGGCGAGTTTATAAAAGAGTGTCTATTGAATGCAGCAGAAATTATGTGTCCCGAACAGAAACAAGCATTTGCAAATGTAAGTCTAACCGGAAATACTGTTGCTCAGCGTGTTGAAGATACGGCTGAGAACTTACAGGACAAGTTGCGTGAAAAAGTTAAATCATTTGTGGCGTTTTCTATTGCAGCTGATGAGAGCACAGATATAAATAATACCACCCAGTTAGCTATATTTATTCGTGGTGTCGATGAGAATTTTGATGTGACCGAAGAACTTTTGGACATGGTGCCCATGACAGGCACAACATCAGGAAATGACTTATTTTTGTGTGTTGAGAAAAGTCTTGAAAAGTTTAATGTAGACTGGTCAAAATTAGTAAGCGTGACTACAGACGGTGCTCCTGCGATGGTCGGTGTTAACGTCGGACTTGTTACGAAACTTAAATCCAAGGTGGCAACGTTTTGCAAGGACACGGAACTTAAGTCCGTTCATTGCATCATTCACCAGGAATCGCTTTGTGCTAAAAAGTTAAAAATGGAACACGTCATGGACGTAGTAATTAACACCGTGAACTGGATACGCTCCCGTGGCTTGAACCACAGACAGTTCAATGCTTTGCTTGATGAATTGGATGCACAATACGGTAGTCTGCTGTACTACACGGAGGTTAGGTGGCTTAGTCGTGGCGTGGTGCTGAAGAGATTTTTTGAACTGTTGAAAGAAATCGACTTGTTCATGTCATCCAAGGGAAAATCCCCGCCTCGGCTCACCAGCGAAGACTGGATCAAAGACTTGGCCTTTTTGGTTGACATTACAACCCATCTAAACACTTTGAATATTTCTCTGCAAGGGCGTTCACAAGTAGTTACACAAATGTATGATTCGATTCGCTCATTCCTAGCGAAATTGTGTCTTTGGGAAACTCATTTGGCAAGGAATAATCTGGCCCACTTTCCTACGCTGAAATCGGTTTCCAGAAATGAAAGTGATGGCCTGAACTACATTCCCAAAATTGTGGAGTTGAAGACTGAATTCCAAAAAAGGTTCTCTGATTTCAAACTTTATGAAAATGAACTAACATTGTTCAGTTCGCCTTTCTCAATTAATATTGATAGTGTGAATGAAGAGCTACAAATGGAAGTTATTGAACTGCAGTGCAATACGGTACTGAAAACTAAATACGACGACGTTGGAATACCAGAATTCTACAAATATCTCGGGAGTAGTTACCCTAAATATAAAAACCATTGTGCAAAGATTCTATCCATGTTCGGAAGTACCTACGTTTGTGAACAGCTGTTTTCCATTATGAAACTGAATAAAACTAAATATCGCTCCCAGTTAAAGGATTCAAGGCTGAATTCTGTACTGCACATCGCAACGCGAAATACAAGACCTGACATTGACATCTTGGTGCAGAAGAAAAGGTGTCAGATTTCTGGTTCCAAATCAAAGGAGTAACGAATTACGAAAGAAAAATTTAATAATTAAATATTTCTATTTTTCAATTTGTATTTTTTCAATTTCGAATTTAAAATATAATCTCCAATATTATACATGTTTGTATTATGTTACATGCTTACGCCATAGTTCAGTAAAAATAAAGTTTGATTATATTTCTGGCAGTTGATTTTTCTTACACCTGGCCCGCTTCACTCATTTACGTTACCTGCCTGGCCCCTGTAGGCATTTGAGTTTGCGACCCCTG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Charlie8 | BPC5 | 407 | 436 | + | -42.85 | AGTGGGAAAACGAAAGATAGACCGAGAGGG |
| Charlie8 | DAL81 | 1326 | 1344 | - | -52.00 | TCAAGCCACGGGAGCGTAT |
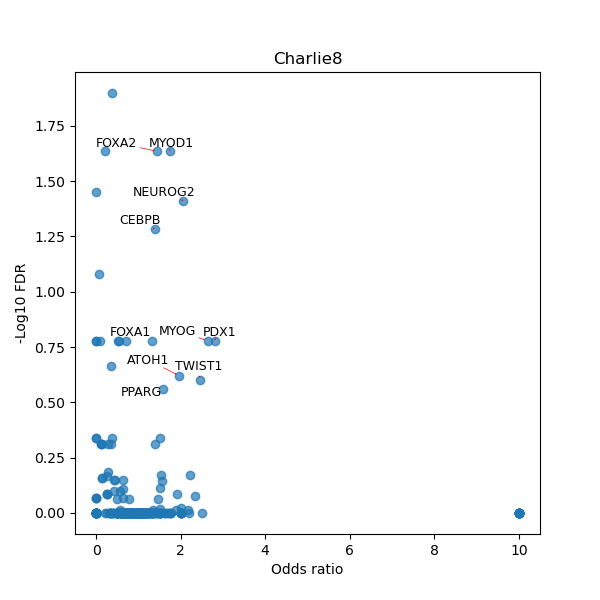
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.