Charlie8
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000107 |
|---|---|
| TE superfamily | Charlie |
| TE class | DNA |
| Species | Eutheria |
| Length | 2457 |
| Kimura value | 28.91 |
| Tau index | 1.0000 |
| Description | hAT-Charlie DNA transposon, Charlie8 subfamily (autonomous) |
| Comment | 15 bp TIR; 8bp TSD with MER1-like NTCTAGAN bias. Nearly complete transposase (roughly <330-2102). |
| Sequence |
CAGAGGTCGCAAACTGGCGGCCCGCGGGCCGNATCCGGCCCGCAGACGTGTTTTGTTTGGCCCGCACAGTGTTTTAAAANATTTTGAATTAGTTGCCAACATTTAAAAATCGGGAGATTTCACATAAAAATCCGGATTTCCGGCTTCTCTTGAAAAATCGGAAGATCTGGCAACACTGGGCCCGCATTCCCGCGTGGCAACAATCGGCTGGAGCTGAGTAGCGGCTGCCCCCTTTAGACGGGGCATGCGCTCTCCAGTTCGCCACAGTCCCCACCACTCCCTATTGTCTCCCCGACACTGAGGCCGAGTGTCAGTTGCCATTTATCATCGCGCTTGCGCTGTTGTTTTTCTTATAGTAGAGTTAAGAGGAAAGTGAAATATTTCTTGTACCCATGTCTCTATCAAAAGTGGGAAAACGAAAGATAGACCGAGAGGGCCGCGTGTTTCAAGAAAAATGGGAGAGAGCATATTTCTTTGTGGAAGTGAAGAATATTCCTACGTGTTTAATATGCAAACAAAGTGTGTCTGTGTCGAAAGAATACAACTTAAGACGCCATTATGAAACAAACCATAGTAAGAACTATGACCGGTATACGGAAAAGATGCGTGATGAAAAACTTAATGAACTGAAAAAAGGACTGAAATTTCAACAGGGTTTGTTTTTGAATGCGAATAAAATAAGTGATGCTGCTGTGAAATGCAGTTATGTATTAAGTGAAAAAATTGCCCGGGCATCAAAACCTTTTACAGATGGCGAGTTTATAAAAGAGTGTCTATTGAATGCAGCAGAAATTATGTGTCCCGAACAGAAACAAGCATTTGCAAATGTAAGTCTAACCGGAAATACTGTTGCTCAGCGTGTTGAAGATACGGCTGAGAACTTACAGGACAAGTTGCGTGAAAAAGTTAAATCATTTGTGGCGTTTTCTATTGCAGCTGATGAGAGCACAGATATAAATAATACCACCCAGTTAGCTATATTTATTCGTGGTGTCGATGAGAATTTTGATGTGACCGAAGAACTTTTGGACATGGTGCCCATGACAGGCACAACATCAGGAAATGACTTATTTTTGTGTGTTGAGAAAAGTCTTGAAAAGTTTAATGTAGACTGGTCAAAATTAGTAAGCGTGACTACAGACGGTGCTCCTGCGATGGTCGGTGTTAACGTCGGACTTGTTACGAAACTTAAATCCAAGGTGGCAACGTTTTGCAAGGACACGGAACTTAAGTCCGTTCATTGCATCATTCACCAGGAATCGCTTTGTGCTAAAAAGTTAAAAATGGAACACGTCATGGACGTAGTAATTAACACCGTGAACTGGATACGCTCCCGTGGCTTGAACCACAGACAGTTCAATGCTTTGCTTGATGAATTGGATGCACAATACGGTAGTCTGCTGTACTACACGGAGGTTAGGTGGCTTAGTCGTGGCGTGGTGCTGAAGAGATTTTTTGAACTGTTGAAAGAAATCGACTTGTTCATGTCATCCAAGGGAAAATCCCCGCCTCGGCTCACCAGCGAAGACTGGATCAAAGACTTGGCCTTTTTGGTTGACATTACAACCCATCTAAACACTTTGAATATTTCTCTGCAAGGGCGTTCACAAGTAGTTACACAAATGTATGATTCGATTCGCTCATTCCTAGCGAAATTGTGTCTTTGGGAAACTCATTTGGCAAGGAATAATCTGGCCCACTTTCCTACGCTGAAATCGGTTTCCAGAAATGAAAGTGATGGCCTGAACTACATTCCCAAAATTGTGGAGTTGAAGACTGAATTCCAAAAAAGGTTCTCTGATTTCAAACTTTATGAAAATGAACTAACATTGTTCAGTTCGCCTTTCTCAATTAATATTGATAGTGTGAATGAAGAGCTACAAATGGAAGTTATTGAACTGCAGTGCAATACGGTACTGAAAACTAAATACGACGACGTTGGAATACCAGAATTCTACAAATATCTCGGGAGTAGTTACCCTAAATATAAAAACCATTGTGCAAAGATTCTATCCATGTTCGGAAGTACCTACGTTTGTGAACAGCTGTTTTCCATTATGAAACTGAATAAAACTAAATATCGCTCCCAGTTAAAGGATTCAAGGCTGAATTCTGTACTGCACATCGCAACGCGAAATACAAGACCTGACATTGACATCTTGGTGCAGAAGAAAAGGTGTCAGATTTCTGGTTCCAAATCAAAGGAGTAACGAATTACGAAAGAAAAATTTAATAATTAAATATTTCTATTTTTCAATTTGTATTTTTTCAATTTCGAATTTAAAATATAATCTCCAATATTATACATGTTTGTATTATGTTACATGCTTACGCCATAGTTCAGTAAAAATAAAGTTTGATTATATTTCTGGCAGTTGATTTTTCTTACACCTGGCCCGCTTCACTCATTTACGTTACCTGCCTGGCCCCTGTAGGCATTTGAGTTTGCGACCCCTG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Charlie8 | Tcf21 | 2041 | 2050 | + | 19.03 | AACAGCTGTT |
| Charlie8 | Tcf21 | 2041 | 2050 | - | 19.03 | AACAGCTGTT |
| Charlie8 | MSC | 2041 | 2050 | + | 18.01 | AACAGCTGTT |
| Charlie8 | MSC | 2041 | 2050 | - | 18.01 | AACAGCTGTT |
| Charlie8 | MYF6 | 2041 | 2050 | + | 17.58 | AACAGCTGTT |
| Charlie8 | MYF6 | 2041 | 2050 | - | 17.58 | AACAGCTGTT |
| Charlie8 | STB3 | 714 | 724 | - | 17.54 | ATTTTTTCACT |
| Charlie8 | TCX2 | 2273 | 2287 | - | 17.51 | ATTTTAAATTCGAAA |
| Charlie8 | ATF4 | 1240 | 1249 | - | 17.50 | ATGATGCAAT |
| Charlie8 | crp | 2040 | 2051 | + | 17.50 | GAACAGCTGTTT |
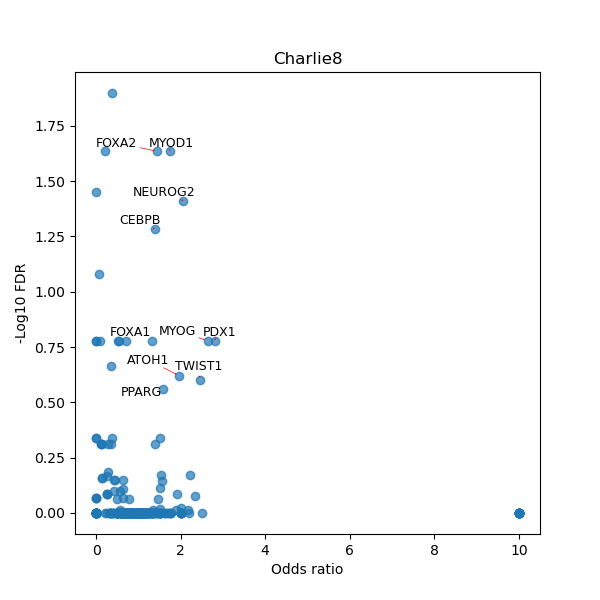
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.