CR1-3_Croc
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001325 |
|---|---|
| TE superfamily | CR1 |
| TE class | LINE |
| Species | Amniota |
| Length | 3605 |
| Kimura value | 33.02 |
| Tau index | 0.0000 |
| Description | CR1 (Chicken Repeat 1) retrotransposon, CR1-3_Croc subfamily |
| Comment | - |
| Sequence |
GGGATCCATCTGACGAGAACGGGGAAGAGCATCTTTGGGCGTCGTCTGGCCAACCTAGTAAGGAGGGCTTTAAACTAGGTTCCACGGGGATGGTGATCAAAACCCTCCAGCAACAGAGCACACAGTCCAGAGGGGAAAGGACTCGAAGAAGCAACTCAACTCTGGGAAAGGGCCAGAGAGCTACAGGACTGCAACAAGTGGCCAAAAGGAAGCGAGTAGGTCAAAATGCAAAATACGTCCGATGCATGTATACAAATGCGAGAAGTATGGGGAATAAACAAGATGAGCTGGAAGCTGCAGCCTGTGAAAAGAACTATGATCTGATTGGCATCACAGAGACCTGGTGGGACAGCTCTTATGACTGGAATGTCAACATCGAGGGTTACACTCTGTTTCGGAAGGACAGGGTTGGGAGAAAAGGTGGTGGTGTTGCCTTGTATGTCAAAAACACATACACTTGTTCCCTGGTTCAGGAGGAAGAAGACAGCAAAGCAGAATGCATTTGGGTGAAGATGAAAGGTGAAAGATGCAGCACCGATATAATGGTGGGGATTTATTATAGGCCACCAAATCAGGAAGAGGAGGTGGATGAGGCACTCTTCAAGCAGGTGACAAATCTATCCAAAAGCTATGGGATGGTACTGATGGGAGACTTCAATTTTCCGGACATCTGCTGGAAGAACAATGCAGCCAAACATAAAATGTCAAGCCGGTTCCTAACTTGTGTGGAGGACAACTTTCTGTTCCAGAAAGTGGAGGATACAACTAGGGGATCATCTATCTTGGATCTTGTCCTGACCAACAGGGAAGAACTGGTGGAGGATGTGAAGGTGATGGGTAACTTGGGAGGAAGCGATCACGATATGATAGAATTCCAGATCCTGAGACGGGGAAAGCATGAGGTCAGCAAAATTAGGACGCTAGATTTCAAAAAGGCGGATTTCATCCGACTCAAGGAGTTAATAGGCATGGTGCCATGGGAGGACAGACTGAAGGATAAAGGCGCCCAGGAGGGCTGGGAGCTTCTAAAGGGCACAGTATTGGAAGCCCAACAAGAAACTGTCCCGACACGATGGAAGCACAAGAAGAATGGCAAGAGACCAATGTGGCTGCACAAGGAGCTTCTAAAATGCCTCAAATGCAAAAGGAAAGCATACAGGCAATGGAAGAATGGGCAGGTCACCAAGGAAGCGTACCAGGAAATAGCAAGAACCTGCAGGGACAAAATCAGGAAGGCAAAGATAAAGAACGAGCTGCACCTGGCGAAGGAGGTTAAGGACAACAAGAAGAGGTTCTATAAGTATGTTGGCCAGAAAAGAAGGACCAAGGAAGCTGTGGGTCCNCTGTTAACAAGCCAGGGGGAACTCCTAACNGAAGATGCAAAGAAGGCGGAGCTACTCAACAGCTACTTTGCCTCAGTCTTCACAAAAAAATTAAGCTGCAGCCAGACACCTAATAAGGATTATCTGGATAAGGAAGGGGGCAAGCTGAGGGANGAGATAACAAAAGAGACGGTCAGAGAGCTTTTGNTGGGTCTGAATGAATTCAAGTCAGCAGGGCCGGATGAACTTCATCCCAGGGTACTGAAGGAACTGGCAGAGGAGATCTCGGAGCCCTTGGCAATGATCTTCATGAAGTCGTGGGAGACGGGCCAGGTACCAGAGGACTGGAAAAGGGCCAACGTAGTGCCCCTCTTTAAAAAAGGCAAAAAGGAGGACCCGGGGAACTACAGACCGGTTAGCCTCACCTCAATACCTGGGAAGTTACTGGAGCATATCCTAAGGAAGGCCATTTGCAAGCATCTGGAGGAGGAAAGGCTGATCACAAGCAGCCAGCATGGATTTACGAAAAACAAATCGTGTCAAACAAACTTGATCTCCTTCTTTGACAAAGTAACTACTTGGGTGGATGAAGGGAATGCTGTGGATATAGTGTATCTGGACTTCAGCAAGGCTTTTGACAAGGTCCCGCATGACCTTCTCATAAACAAATTGGAGAAATGTGGGCTGGACAAAACTACTACAAGGTGGATAGATAACTGGCTCAATAGCCGCAAGCAAAGGGTAATTATTAACGGGTCCATGTCAGAATGGCAGGAGGTTTCGAGTGGAGTCCCACAGGGGTCTGTCCTGGGCCCGGTGCTGTTCAACGTGTTTATTAATGATTTGGATGCTGGCATAGAAAGCTTCTTGAGCAAGTTTGCAGACGACACGAAACTGGGAAGGATTGCGAACACGTTGGAGGACGNGAGCGCAATTCAGAAGGATCTCGACAGATTGGAAAGCTGGGCTAGAGCTAACAGGATGAAGTTCAATGCGGACAAATGCAAGGTGCTACACCTCGGGAGGAATAATCAAAAACACAAATACAAAATGGGTGACACCTGGCTGGATGGCAGCACTATGGAAAAGGACCTGGGAGTCTTGGTAGATCACAGACTCAATATGAGTCTGCAGTGTGATGCAGCCGCACAAAAGGCGAATGCGGTTTTGGGATGCATCAATAGAAGCATTAGGTGCAAGACACGGGAGGTGATAGTGCCTCTCTACTCGGCACTGGTTAGGCCTCATCTGGAGTACTGTGTGCAGTTCTGGGCTCCACATTTTAAAAAGGACGTGGAGAAGTTAGAGAGGGTCCAGAGGCGGGCAACAAAGATGATAAAGGGCTTGGAAGGCAAGTCATACGAGGAGAGGCTGAAGGAGCTAGGCATGTTCAGCTTGCGGAAAAGGCGCTTAAGAGGGGACATGATAGCAGTCTTCAAATACTTGAAGGGCTGCCATAAAGAAGAGGGAAAGCACCTTTTCTCTCTTGCTGCAGAGAGGAGGACGCGGACCAATGGCTTGAAGTTGCAGCAAAGTAGGTTTAGATTGGATATCCGGAAAAACTTCTTCACTGTTAGAACAGTGAGGCAGTGGAATAGACTGCCTAGGGAAGTTGTGGACTCTCCATCACTGGAGGTGTTCAAGAAGAGGTTGGACAGCCACTGGTCGGGGATGATCTAGGCGTAGCTATGCCTATGTTCTACTATGGGTATTTCCCATGCTTCTGGGCTTTGCTGGTTGCCCCCGCCTCCCCCCCCCCCCCTTTCTGCTACATGTGTCAGGGTGTTTTTAAGCCTTCCTCAGAGGCATTGGGTGTTGGCTGCAGCCAAGGCTGGGGACTTTGACTGGGGTGTGCCAATGCTCCTGCTGGGACCAAAGCCGGAGGTTCCTGGCTAGAGGGTCTTGCCCCTCCGCTCAGGGTCAGACCGATCGCCATATTTGGGGTCAGGAAGGAATTTTACCCCATGGTCAGATTGGTACGGACTGTGGGGGTTTTTGCCTTCCTCTGTAGCGGGGGGCGTGGCCCTCTACCCGGGATCTCTTGAGCATATATTGACAACATTTCATAGAAGCAGGACATTGGCTGCCGTGGTTCCCCTGCTTTACCTGTGGCAGGTTAGGGTGTTAGGTCTGTGTTGTGTCAGGGTTGACGGTAGTCTTACGTAGAGTTTAGATTATGGTTGTATGGGATGGTTTGGATAGGGATGATCCTGCCTCAGGCAGGGGGTTGGACTAGATGACCTCTGGAGGTCCCTTCCAGCCCTACTTCTCTATGATTCTATG
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| CR1-3_Croc | ZNF460 | 3062 | 3077 | + | 17.55 | GCCCCCGCCTCCCCCC |
| CR1-3_Croc | NAC075 | 2182 | 2201 | + | 17.44 | AAAGCTTCTTGAGCAAGTTT |
| CR1-3_Croc | PATZ1 | 3063 | 3073 | - | 17.39 | GGAGGCGGGGG |
| CR1-3_Croc | NAC005 | 2183 | 2201 | + | 17.34 | AAGCTTCTTGAGCAAGTTT |
| CR1-3_Croc | Klf6-7-like | 3064 | 3083 | + | 17.30 | CCCCGCCTCCCCCCCCCCCC |
| CR1-3_Croc | Klf15 | 3065 | 3086 | + | 17.24 | CCCGCCTCCCCCCCCCCCCCTT |
| CR1-3_Croc | DOF4.2 | 1699 | 1709 | + | 17.21 | AAAAAGGCAAA |
| CR1-3_Croc | SP9 | 3336 | 3345 | - | 17.16 | CCACGCCCCC |
| CR1-3_Croc | KLF13 | 3331 | 3347 | - | 16.64 | GGCCACGCCCCCCGCTA |
| CR1-3_Croc | scrt | 191 | 201 | + | 16.61 | GCAACAAGTGG |
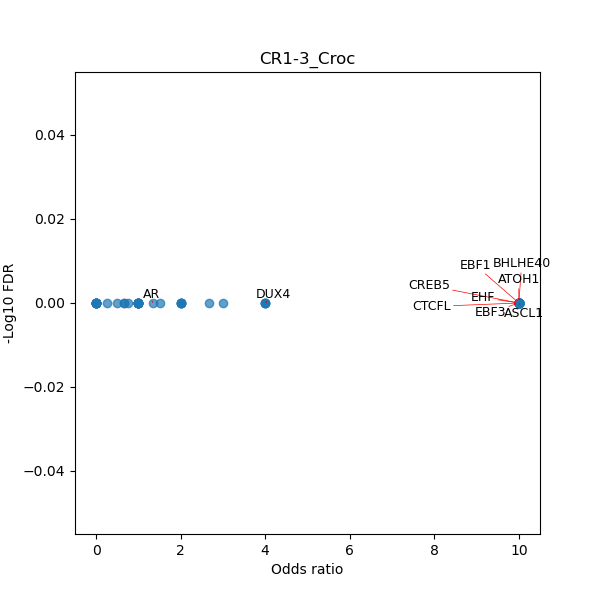
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.