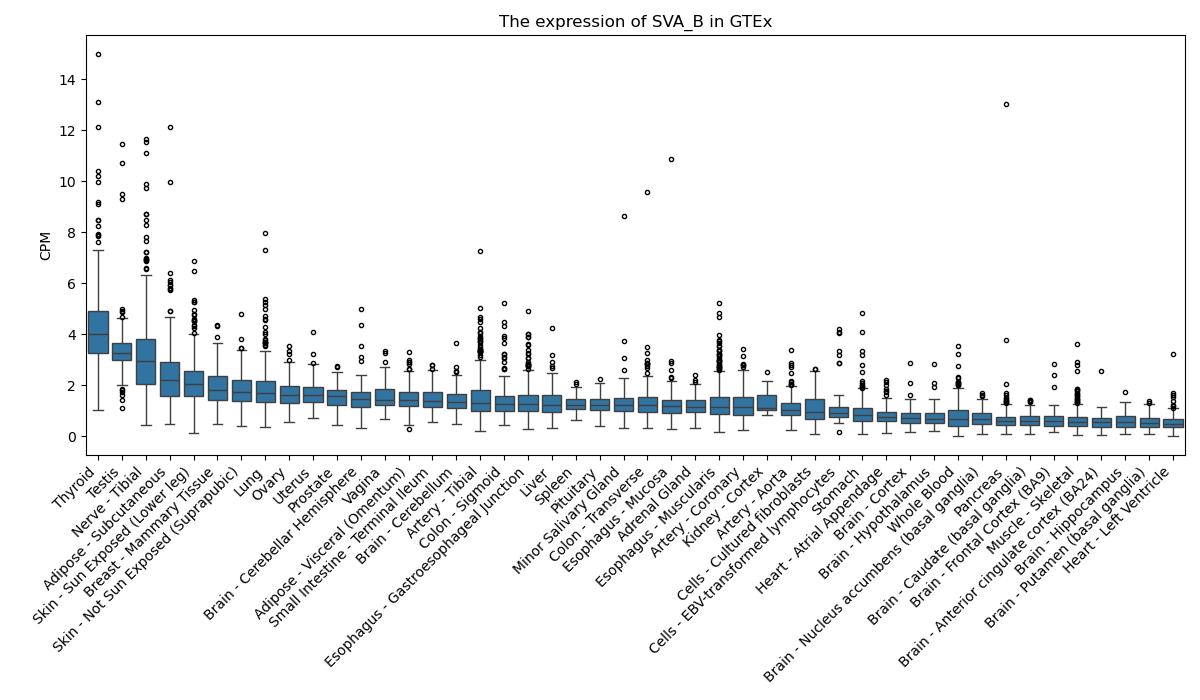
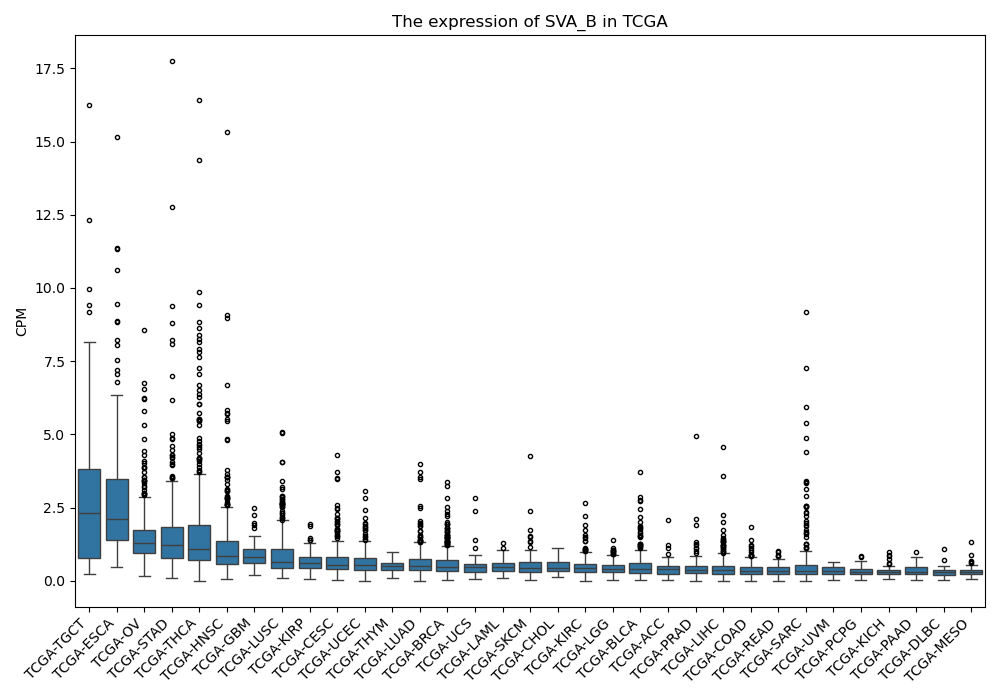
SVA_B
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001068 |
|---|---|
| TE superfamily | SVA |
| TE class | Retroposon |
| Species | Hominidae |
| Length | 1383 |
| Kimura value | 5.80 |
| Tau index | 0.6953 |
| Description | Composite retroelement: (SINE-R + VNTR + Alu), SVA_B subfamily |
| Comment | SVA retroelements are hominid-specific chimeras, some of which are still active in the human population. These SINEs are a unique combination of features. The 5' end begins with hexamer repeats, followed by two Alus in reverse orientation. The center is comprised of a number of VNTR regions (Variable Number of Tandem Repeats) that may span a few hundred basepairs, up to 2000bp. The 3' end is formed by a portion of a HERVK LTR (the SINE-R feature). |
| Sequence |
CTCTCCCTCTCCCTCTCCCTCTCCCTCTCCCTCTCCCTCTCCCTNTCCCCTCTTTCCACGGTCTCCCTCTGATGCCGAGCCGAGGCTGGACTGTACTGCCGCCATCTCGGCTCACTGCAACCTCCCTGCCTGATTCTCCTGCCTCAGCCTGCCGAGTGCCTGGGATTGCAGGCGCGCGCCGCCACGCCTGACTGGTTTTCGTATTTTTTGGTGGAGACGGGGTTTCGCCGTGTTGGCCGGGCTGGTCTCCAGCTCCTGACCGCGAGTGATCTGCCNGCCTCGGCCTCCCGAGGTGCCGGGATTGCAGACGGAGTCTCGCTCACTCAGTGCTCAATGTTGCCCAGGCTGGAGTGCAGTGGCGTGATCTCGGCTCGCTACAACCTCCACCTCCCAGCCGCCTGCCTTGGCCTCCCAAAGTGCCGAGATTGCAGCCTCTGCCCGGCCGCCACCCCGTCTGGGAAGTGAGGAGCGTCTCTGCCTGGCCGCCCATCGTCTGGGATGTGAGGAGCCCCTCTGCCCGGCCGCCCAGTCTGGGAAGTGAGGAGCGCCTCTTCCCGGCCGCCATCCCGTCTGGGAAGTGAGGAGCGTCTCTGCCCGGCCGCCCATCGTCTGGGATGTGGGGAGCGCCTCTGCCCCGCCGCCCCGTCTGGGANGTGAGGAGCGCCTCTGCCCGGCCAGCCGCCCCGTCTGGGAGGTGAGGAGGTCAGCCCCCCGCCCGGCCAGCCGCCCCGTCCGGGAGGAGGTGGGGGGNNCAGCCCCCCGCCCGGCCAGCCGCCCCGTCCGGGAGGTGGGGGGCGCCTCTGCCCGGCCGCCCCGTCTGGGAAGTGAGGAGCCCCTCTGCCCGGCCGCCACCCCGTCTGGGAGGTGTACCCAACAGCTCATTGAGAACGGGCCATGATGACGATGGCGGTTTTGTCGAATAGAAAAGGGGGAAATGTGGGGAAAAGAAAGAGAGATCAGATTGTTACTGTGTCTGTGTAGAAAGAAGTAGACATAGGAGACTCCATTTTGTTCTGTACTAAGAAAAATTCTTCTGCCTTGGGATGCTGTTAATCTATAACCTTACCCCCAACCCCGTGCTCTCTGAAACATGTGCTGTGTCCACTCAGGGTTAAATGGATTAAGGGCGGTGCAAGATGTGCTTTGTTAAACAGATGCTTGAAGGCAGCATGCTCGTTAAGAGTCATCACCACTCCCTAATCTCAAGTACCCAGGGACACAAACACTGCGGAAGGCCGCAGGGTCCTCTGCCTAGGAAAACCAGAGACCCTTGTTCACATGTTTATCTGCTGACCTTCCCTCCACTATTGTCCTATGACCCTGCCAAATCCCCCTCTCCGAGAAACACCCAAGAATGATCAATAAATACTAAAAAAAAAAAAA
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| SVA_B | BPC5 | 3 | 32 | - | 17.32 | AGGGAGAGGGAGAGGGAGAGGGAGAGGGAG |
| SVA_B | BPC5 | 9 | 38 | - | 17.32 | AGGGAGAGGGAGAGGGAGAGGGAGAGGGAG |
| SVA_B | MYB88 | 464 | 472 | - | 17.21 | ACGCTCCTC |
| SVA_B | MYB88 | 580 | 588 | - | 17.21 | ACGCTCCTC |
| SVA_B | PATZ1 | 709 | 719 | - | 17.07 | CCGGGCGGGGG |
| SVA_B | PATZ1 | 757 | 767 | - | 17.07 | CCGGGCGGGGG |
| SVA_B | ZNF460 | 401 | 416 | + | 16.88 | GCCTTGGCCTCCCAAA |
| SVA_B | IG1 | 96 | 104 | + | 16.76 | CTGCCGCCA |
| SVA_B | ZNF460 | 380 | 395 | + | 16.56 | ACCTCCACCTCCCAGC |
| SVA_B | Zm00001d002364 | 633 | 642 | + | 16.53 | CCCCGCCGCC |
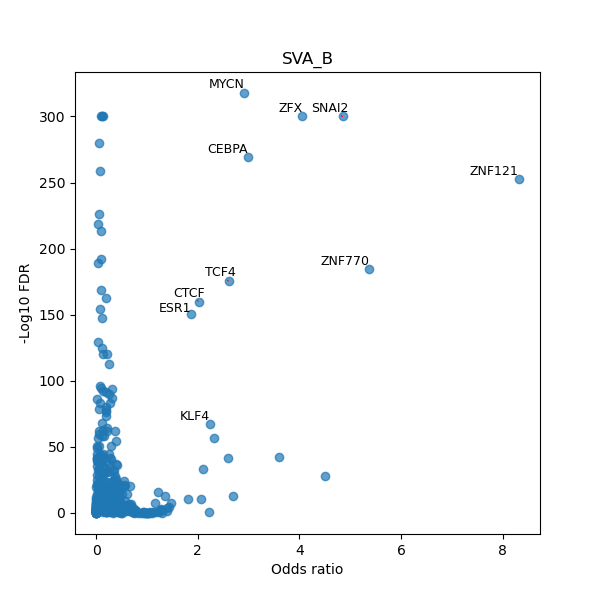
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.