L1MEd_5end
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0000777 |
|---|---|
| TE superfamily | L1 |
| TE class | LINE |
| Species | Eutheria |
| Length | 2224 |
| Kimura value | 31.29 |
| Tau index | 0.9542 |
| Description | 5' end of L1 retrotransposon, L1MEd_5end subfamily |
| Comment | 5' end of LINE elements with L1ME subfamily 3' ends ORF1 starts at pos. 682. ORF 684-1640 encodes a gag protein similar to those of other L1ME elements. ORF2 starts at position 2075 (standard first 150 nucleotides included). |
| Sequence |
CGGACTTCCGTTTCCGGCAATATGGCGGACTAGATAACCTGAAAACCCTCCCGCTACAAAACACCTAGAAATGCTGGATAAAATATAACAAACATCCTTTTAAATGCATAGCTGAGCTCGCAAGAAAGTAAGGGAAATCCCCAGGGGCCAAAAACGAAGAGGGAACTGAAAACCAGAGCGGTAAGCACGAGCTGAAGCTGCGGCTGCCCTGAGGGCATTTGCCGGTCTCGGTAACCNAGGGGCTTGGGTTTTAACGGCCACGCGGGGACAGGAGACGAGGCCTTGGGCCCGCGCAAGGCGGGGAGTTGGAACTGAGACCCCCGCATAAAGCCGGGACCCTCGAAGGGCTACACCCTCAGTGAAAGGGTGGACTAGAAAAAAATCCGCCCACCGGCACAGGGAGACGACAAGGAAACTTGTCTGTCTCGGCCTGGGCTCTGGGTGGAGAAAAAAAGTCTCCCCTGAGAATTCGTAACCACAGGCCTGCCCTCACGCGGGTTTGGGGTTCGAATTTACACTACCTGCGTGGTCCGGGAAACCCCAAGCCGAGAAATTAACTTAAAGTGGTCCCGGGCTGGTAGTACCCCTGGGGCGCCTGGCAGAAGCAAACGCAAANCCTCTCTGGAGGAACGCACCCTCAACCCAGGCCNCANAGGATTCCCACAGATAAAGCCCCGNCGAANATGAGCTCACAATCNAAAATTACAAAACACACGAGGAAACAANCCACCATGAGCGAGAGTCAGCAGAAACAACAAACAGCAGAATTAGACCCCCAAGAACTTCAGATANTGGAATTATCAGATANAGANTATAAAATAAGTATGTTTAAAATGNTTAAAGANATAAAAGAAGGAATTAAAAACATGAGNAAGGAACAAGANACTATNAAAAANGACCAGGCAGATTTGAAAAAGAACCAAATAGAACTTCTAGAAATGAAAAATATAGTCATTGAAATTAAAAACTCAATGGATGGGTTAAACAGCAGATTAGACACAGCTGAAGAGAGAATTAGTGAACTGGAAGATAGATCTGAAGAAATTACCCAGAATGCAGCACAGAGAGATAAAGAGATGGAAAATATGAAAGAGAGGTTAAGAGACATGGAGGATAGAATGAGAAGGTCTAACATACGTCTAATNGGAGTTCCAGAAGGAGAGAATAGAGAGAATGGGGGAGAGGCAATATTCGAAGAGATAATGGCTGAGAATTTTCCAGAATTGATGAAAGACATNAATCCTCAGATTCAGGAAGCNCAACGAATCCCAAGCAGGATAAATAAAAANAAATCCACACCTAGACACATCGTAGTGAAACTGCAGAACACCAAAGACAAAGAGAAGATCTTAAAAGCAGCCAGAGAGAAAAGACAGATTACCTACAAAGGAACGACAATTAGACTGACAGCNGACTTCTCAACAGCAACAATGGAAGCCAGAAGACAGTGGAATAATATCTTCAAAGTGCTGAGAGAAAATAACTGTCAACCTAGAATTCTATACCCAGCNAAACTATCNTTCAAGAATGAGGGCGAAATAAAGACATTTTCAGACAAACAAAAACTGAGAGAGTTTACCACCAACAGACCCTCACTAAAGGAACTNCTAAAGGATGTACTTCAGGAAGAAGGAAANTGANCCCAGAAGGAAGGNCTGAGATGCAAGAAGGAATGGTGAGCAAAGAAANTGGTAAACATGTGGGTAAATCTAAACAAACATTGACTGTATAAAACAATAATAATAATGNCTAATTTGNGGGGTATAAAAACAAGGTAGAACTAAAATACTGGACAACAATAACATGTAAGTCGGGAGGGGGGTGATCGGAGTTAAAGCGTTCTAAGGTCCTTGTATTGTTCGGGAGGAGGGTAAAGATATTGATTAACTTTAGACTTTGTTAAGTTAAGTATGCATGTTAAAATTTTAAGGGTAACCACTAAAAGAATAGAAATAGAATGTATAACTTCCAAACCAGTAGAGGGGAAAANAAAAAANAAAAANNNAATCAATCCAAAAGAAGGCAAGAAAGGAGAGAAAAAGAAACATAGAACAAATAGAAAGCACAAAATAAGATGGTAGAAATAAATCCAAATATATCAGTAATCACAATAAATGTAAATGGACTAAACTCNCCAGTTAAAAGACAGAGATTGTCAGATTGGATTAAAAAAACAAATCCAGCTATATGCTGTTTACAAGAGACACACCTAAAACATAAGGAC
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| L1MEd_5end | NAC071 | 1618 | 1632 | - | 16.77 | CTTCTTCCTGAAGTA |
| L1MEd_5end | FOXD2 | 1710 | 1720 | + | 16.58 | CTAAACAAACA |
| L1MEd_5end | ARALYDRAFT_496250 | 564 | 571 | - | 16.55 | GGGACCAC |
| L1MEd_5end | FOXE1 | 2166 | 2177 | + | 16.40 | TTAAAAAAACAA |
| L1MEd_5end | F10B5.3 | 1214 | 1224 | + | 16.35 | TTTTCCAGAAT |
| L1MEd_5end | DOF5.1 | 2037 | 2055 | + | 16.27 | AAAAAGAAACATAGAACAA |
| L1MEd_5end | TCP3 | 564 | 571 | - | 16.23 | GGGACCAC |
| L1MEd_5end | TCP24 | 564 | 571 | - | 16.21 | GGGACCAC |
| L1MEd_5end | NFkb | 132 | 142 | + | 16.19 | GGGAAATCCCC |
| L1MEd_5end | TCP5 | 564 | 571 | - | 16.15 | GGGACCAC |
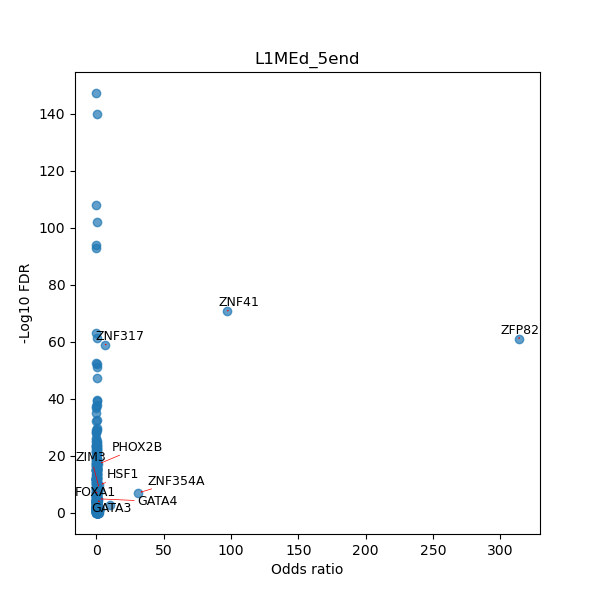
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.