Eutr5
Basic information Differential Expression Stage analysis Survival analysis Correlation analysis| DF ID | DF0001258 |
|---|---|
| TE superfamily | Long_Terminal_Repeat_Element |
| TE class | LTR |
| Species | Eutheria |
| Length | 1130 |
| Kimura value | 34.16 |
| Tau index | 0.9620 |
| Description | Ancient interspersed repetitive sequence from eutherian mammals. |
| Comment | >71% identical to consensus. Origin unknown. More than 200 copies in eutherian mammals. Not present in opossum. ; but it shows no similarity to known SINEs and LINEs. Provisionally guessed LTR because of TG...CA termini but no obvious TSDs nor an obvious (AWTAAA) poly A signal in either orientation. GA-rich 5' end and C-rich satellite-like 3' tail support classification as a SINE. Present at orthologous sites in elephants and humans, absent in opossum. |
| Sequence |
TGTGGCGGGATGAACTGAGATCCTCTTATCCATTCCCCTCCCTCCCGCGGTCAGGGGAGAAACCTGGGGATTAGAGAGAAGCGATTTGGGCATGGCGCCATTCATTGTCAGTTAATGTGAGGAAGCTTTGTAATGTAACCCAGGGCAGGGAAGGGGCAAATGCTCTCTTAGCATAATAAGGCCAGGGTTATTATTTCGCAAGAAATAGGGTCAAAGTTATGTTAATAGGAGGAATATGAATTTCTCATGTAGACCAGACTGGGTGGTCTTCGGAAGGAAATTAATCCTCGCGGTGGCGAGGTGTGTTTGATTTTAGATACTTTAGGAGAAAACCAGCCGCAGCCCGGAGTGGAAGCCCCACTCTGTTCCCACCCCCCTCCCCTNCCCCCAGGACCCAGAGCTCACCCGCCTCCTTTCCTGGGGAGCCAGCGGACCCTGAGACCAGAGCCCACCCCTCCTCTGCAGAGCCAGCAGAANCTGGGCAGAGGAGAAAAGCCACTGACATTCAGCGGACGCCTGAGGCTGCGGAATTTGCAAGACACAAGAAGGGACTAGGGAGACTGAGACTTGATTATTTCTGTAGGGAAGGGTGGCGGATCCCGGGAGAAAGAGGGATTGGATGGGGAAGCAGAAGGAGAAGAGGTTAAAGGGAGAGGGCNGNAGAAATGAGGGAGAAGATAGGAAGCGAGGGAGCGTCCACGTGGGACGCCCTCCGAAGGCCTAGGCAGTGAGGCCTCGTCCCTTGTGTGGTACAAGGGAGCGAGGTAGCTAGACCAGAGNCGCAATCTCCCATCTCCTTCCTCCATCCCGCTTTGGGACAATTACTTGAACCCTTCAGTAAAGAACTGCTTAAACCTCCATCGGAGTCCAGCGTGTATTCGGTTCGGGGAAGACGGGCGAGGCGGTATTCGGANCGGGGAGGGATAACCCCGCTCCNCGGCCATCTGATGGGCATCGGCAGCNCCCTCNCTGTGGACTACTGGGCGGCGAGGAGCATCNACCCGCAGACGACGACCCAGCTGGAGAAACGGTAAGGCCGATCCCCCCGACTTCCCTCCCCCCAGAGAGGCGACTAGAGACATNATCTGTGGGACAGAAAAGGGAGGAGCCCGCCACATGGGTCACATACA
|
TF motifs of the concenus sequence
Use FIMO to detect transcription factor motifs in the concenus sequence of the TE family.
| TE_family | TFBS | Start | End | Strand | Score | Matched sequence |
|---|---|---|---|---|---|---|
| Eutr5 | TFAP2B | 515 | 523 | + | 15.56 | GCCTGAGGC |
| Eutr5 | TfAP-2 | 515 | 523 | - | 15.55 | GCCTCAGGC |
| Eutr5 | BHLH72 | 697 | 705 | - | 15.53 | CCCACGTGG |
| Eutr5 | NFIX | 87 | 100 | + | 15.51 | TGGGCATGGCGCCA |
| Eutr5 | ZNF281 | 1052 | 1061 | - | 15.49 | GGGGGAGGGA |
| Eutr5 | MYCN | 697 | 704 | + | 15.35 | CCACGTGG |
| Eutr5 | MYCN | 697 | 704 | - | 15.35 | CCACGTGG |
| Eutr5 | RARA::RXRG | 142 | 158 | + | 15.28 | AGGGCAGGGAAGGGGCA |
| Eutr5 | ZNF768 | 1062 | 1070 | - | 15.25 | GCCTCTCTG |
| Eutr5 | HINFP | 509 | 516 | - | 14.82 | GCGTCCGC |
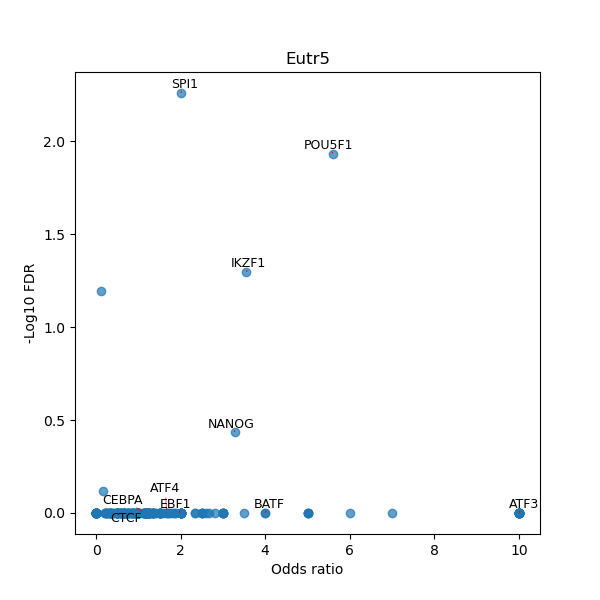
TFBS enrichment in GRCh38
Use Fisher's exact test to perform enrichment analysis of transcription factor binding sites in the TE family of GRCh38.